320k Human CRISPRi K562 cells, Multiplexed Samples, 16 Barcodes, Ultima Sequencing
Flex dataset analyzed using Cell Ranger 9.0.1
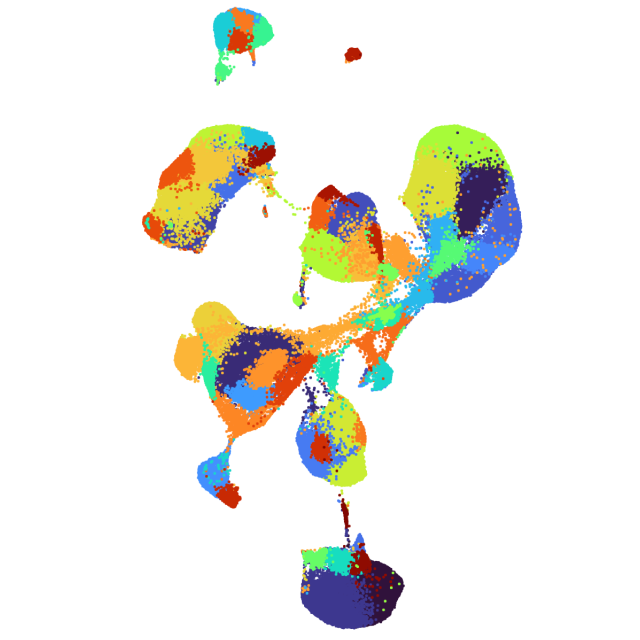
Learn about Chromium Analysis
Biomaterials
A human K562 cell line, stably expressing KRAB-dCas9, was transduced with a pooled lentiviral CRISPRi lncRNA and protein coding library containing approximately 6,903 sgRNAs. The CRISPRi library consisted of 4,045 sgRNAs, including those targeting 280 lncRNAs, 567 protein-coding genes, and 177 non-targeting sgRNA controls. The target genes for the remaining 2,681 sgRNAs were unknown and set to "ignore". Transduced cells were allowed to recover for 48 hours, followed by four days of culture in selection media. The CRISPRi screen was stopped on Day 6 and the selected cells were cryopreserved.
Sample preparation
Cryopreserved cells were thawed, counted, and fixed for 24 hours at 4°C following the demonstrated protocol Fixation of Cells & Nuclei for GEM-X Flex Gene Expression (CG000782).
Assay workflow
Gene Expression and matching CRISPR libraries were generated as described in the GEM-X Flex Gene Expression Reagent Kit for Multiplex samples (CG000787) with the workflow modifications described in the Knowledge Base article, What workflow modifications are required when performing CRISPR Screening with the Single Cell Gene Expression Flex assay?
The fixed cells were run as a 16-sample multiplexed experiment, where the sample was evenly split across 16 separate hybridization reactions, each using a unique Probe Barcode (sample sub-pooling). Custom probes designed to capture the sgRNAs and their gene targets were pooled and spiked in per the Custom Probe Design for Visium Spatial Gene Expression and Chromium Single Cell Gene Expression Flex Technical Note (CG000621). After hybridization, cells from each hybridization were pooled in equal proportions, washed, and then run on a GEM-X Flex chip.
Sequencing
Libraries were sequenced on an Ultima UG 100™ sequencer following the UG 100 Solaris™ Flex workflow to 8,789 reads per cell for the gene expression library and 8,654 reads per cell for the CRISPR library.
Analysis
Cell Ranger v9.0.1 was used to map reads to the probe set and output feature-barcode matrices for further analysis.
How to view data
To get started, download Loupe Browser to explore the Loupe file, or read more about the other Cell Ranger outputs specific to Flex data.
This dataset is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) license. 10x citation guidelines available here.
