Preview Data: Atera In Situ Gene Expression, FFPE Human Cervical Cancer
Atera dataset analyzed using Atera Onboard Analysis dev
Learn about Atera
Overview
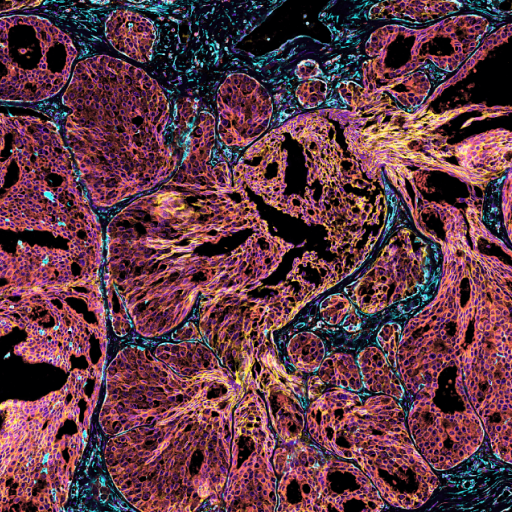
This human FFPE cervical cancer data showcases results using the pre-commercial version of the Atera Whole Transcriptome Assay (WTA), which is currently under development. The assay was designed to closely match the Chromium Flex Apex assay in terms of content and sensitivity, and includes 18,028 genes. The number of probes per gene were optimized to maximize sensitivity of lowly expressed genes and avoid optical crowding risk for highly expressed genes. The 10x Genomics in situ multimodal cell segmentation solution was used to generate cell segmentation boundaries.
This sample is from the same block as in this Xenium Prime 5K dataset and this Single Cell Gene Expression Flex dataset.
How to view data
Interactively explore this dataset with the web demo ("Open web demo" above), a preview of the upcoming desktop version of visualization software.
This preview dataset can also be opened in Xenium Explorer software (v4.1.1), but you may experience performance issues due to the large dataset size. See the Getting Started with Xenium Explorer page for general guidance navigating the web demo interface and these instructions to explore the aligned H&E image in the demo.
Download the files to explore further on your desktop. This preview dataset was converted to closely resemble Xenium Onboard Analysis v4 file formats for compatibility with existing analysis packages. It has been tested with seurat v5.4.0, scanpy v1.12.0, and spatialdata v0.7.2 + spatialdata_io v0.6.0. However due to minor differences in formats, it is not compatible with Xenium Ranger and not guaranteed to be compatible with other untested community-developed tools. Atera's official output data format will be optimized to integrate with the broader bioinformatics ecosystem and reduce data management overhead. Future Atera data releases will showcase the new file format to help customers transition.
Biomaterials
FFPE-preserved tissue blocks were purchased from Discovery Life Sciences (squamous cell carcinoma: III-B (T1b N1 MX)).
Analysis metrics
| Metric | Cervical Cancer |
|---|---|
| Median transcripts per cell | 929 |
| Cells detected | 717,576 |
| Nuclear transcripts per 100 µm² | 2,252.0 |
| Total high quality decoded transcripts | 1,040,030,708 |
| Region area (µm²) | 79,418,423.2 |
Custom Cell Groups
Differentially expressed genes from the graph-based clustering results were exported to annotate cell types. Major cell groups were annotated using published cervical cancer cell atlas references. T cell subtypes were identified using known marker genes. The endocervix was identified based on the spatial location and expression of mucins. Well-differentiated tumor subtypes of the ectocervix (basal, parabasal, dyskeratotic) were annotated based on their spatial location in the columnar epithelium.
This dataset is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) license. 10x citation guidelines available here.
