Human Breast Cancer: Whole Transcriptome Analysis
Spatial Gene Expression dataset analyzed using Space Ranger 1.2.0
Learn about Visium Analysis
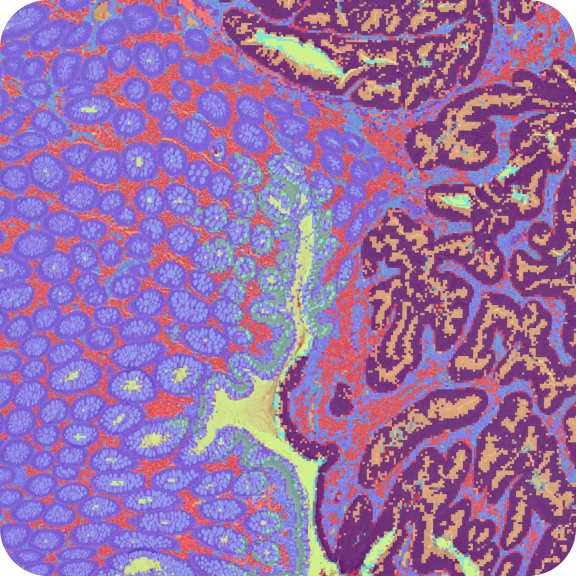
10X Genomics obtained fresh frozen human Invasive Lobular Carcinoma breast tissue from BioIVT Asterand. The tissue was embedded and cryosectioned as described in Visium Spatial Protocols – Tissue Preparation Guide Demonstrated Protocol (CG000240). Tissue sections of 10µm were placed on Visium Gene Expression slides and fixed and stained following Methanol Fixation, H&E Staining & Imaging for Visium Spatial Protocols (CG000160).
The tissue was AJCC/UICC Stage Group I, ER positive, PR positive, HER2 negative.
The H&E image was acquired using a Nikon Eclipse Ti2-E microscope with the following settings:
- Color camera
- 10X objective
- Numerical Aperture: 0.45
- Exposure: 20 ms
The Visium Gene Expression library (T1T2-A6) was prepared as described in the Visium Spatial Reagent Kits User Guide (CG000239) Rev D). Sequencing data was processed using Space Ranger.
- Sequencing instrument: Illumina NovaSeq 6000, flow cell HHYWHDSXY (lanes 1-4)
- Sequencing depth: 72,436 mean reads per cell
- Sequencing configuration: Paired-end (28 X 90), Dual-Indexed Sequencing. Read 1: 28 cycles (16 bp barcode, 12 bp UMI); i7 index: 10 cycles; i5 index: 10 cycles; Read 2: 90 cycles (transcript).
- Slide: V19L29-095
- Area: A1
Key metrics were:
- Spots detected: 4,325
- Median genes per spot: 3,671
- Median UMI counts per spot: 9,465
To maintain donor anonymity, sequence data for this sample is not currently available.
This dataset is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) license. 10x citation guidelines available here.
