Delving deeper into transcriptomic changes in non-neuronal brain cells in Alzheimer’s disease
While we have gleaned substantial insight into the development and progression of Alzheimer’s disease (AD)—namely, the existence of hallmark amyloid-beta protein deposits and neurofibrillary tangles—we still lack a deeper understanding of what cell-by-cell changes drive the disorder. At this year’s 10x Genomics Global Neurodegenerative Disease Summit, held virtually August 23–26, Nancy Ip, PhD, of the Hong Kong University of Science and Technology will present recent work (1) using single cell transcriptomic analysis to reveal how non-neuronal cell types drive AD progression, including endothelial cell angiogenesis.
AD is the most common type of dementia, affecting 24 million people around the world—a rate that is expected to double every 20 years (2). For patients and families alike, the fact that there is no cure and few effective treatments—currently medications only treat the earliest stages of cognitive decline—makes AD one of the more devastating diseases of aging.
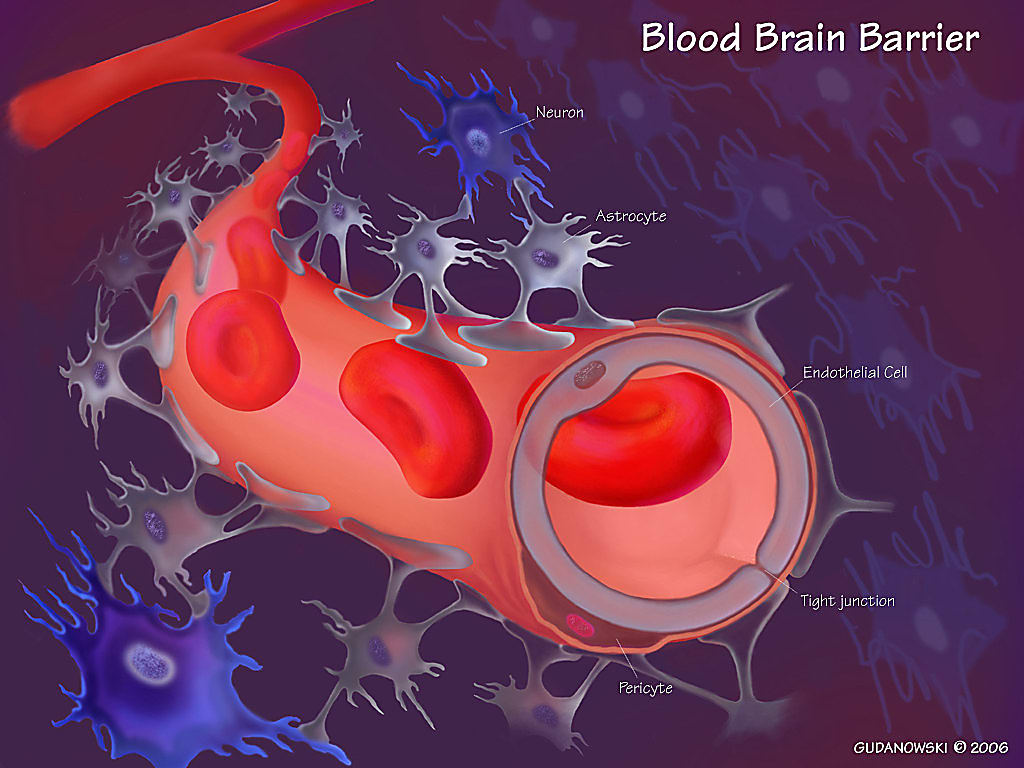
In an effort to unravel the multicellular contribution of brain cells, immune cells, and environmental factors, scientists have turned to single cell technologies. During AD, many changes take place to cause degeneration of multiple cell types and dysfunction of signaling pathways, including altered synaptic function (3), a decrease in blood–brain barrier integrity, and the loss of neurotrophic support (4). Previous studies used single cell transcriptomic analysis to profile changes in AD neurons and oligodendrocytes, which are more predominant brain cell types (5–7). However, studies of transcriptional changes and molecular pathways in other cell types, specifically endothelial cells, which line the blood–brain barrier and help maintain its structure, have been lacking.
Cell type–specific transcriptomic profiling reveals complex roles for glial cells
Dr. Ip and team used Chromium Single Cell Gene Expression to perform single nucleus RNA-sequencing (snRNA-seq) on 169,496 nuclei from prefrontal cortex tissue samples of AD patients and healthy controls. They identified six main cell types, including glial cells (which maintain brain homeostasis in various ways), excitatory and inhibitory neurons, and endothelial cells.
Among the 2,190 differentially expressed genes (DEGs), only 11 were differentially expressed in all cell types, suggesting that expression changes in AD are mostly specific to each cell type. From these DEGs, they found that AD-related, cell type–specific transcriptomic changes in astrocytes are associated with synaptic signaling, microglia with immune response, oligodendrocytes with myelination, and endothelial cells with both angiogenesis and immune response.
Teasing apart subpopulations of cells within astrocytes, microglia and oligodendrocytes lent even further insight. In AD samples, an astrocyte subpopulation important to neurotransmitter recycling was reduced while a population of astrocytes expressing alarmin response genes were induced, suggesting that astrocytic dysfunction contributes to impaired neurotransmitter recycling. For microglia, a subpopulation that expresses genes important for synaptic pruning was reduced, suggesting that the loss of these cells can contribute to altered synaptic pruning. Finally, they saw a decreased number of mature myelinated oligodendrocytes and an increase in the number of remyelinating oligodendrocytes, suggesting demyelination may be a key event in AD.
Angiogenic endothelial cells help drive immune dysfunction and progression in AD
Endothelial cells maintain the blood–brain barrier and thus are important mediators of the interaction between the immune system and brain cells. Dr. Ip’s profiling analysis showed that transcriptomic changes in endothelial cells in AD were associated with both angiogenesis and the immune response. Specifically, her team identified three AD-increased subpopulations of endothelial cells that were enriched for genes associated with angiogenesis and antigen presentation—especially major histocompatibility complex class I molecules. Signature genes of these cell types form an interaction network associated with angiogenesis/adhesion, transmembrane transport, antigen presentation, metal ion homeostasis, cellular respiration, and rRNA processing. Using these genes, ingenuity pathway analysis revealed that a proinflammatory response, conserved between mice and humans, induced angiogenic endothelial cells.
Toward a nuanced understanding of cell type–specific changes in AD
In summary, single cell transcriptomic profiling is a useful tool to map gene expression changes occurring not only in different brain cell types, whether neuronal, glial, or endothelial, but also in subpopulations within these types. Increasing our understanding of the mechanisms underlying the how and why different brain cell types contribute to AD progression will allow scientists to discover new molecular targets for treating this debilitating and often fatal neurodegenerative disease.
We believe generating actionable insights that will improve the lives of patients with neurodegenerative diseases, and one day prevent these diseases altogether, requires a holistic view of these complex pathologies. By hosting the 10x Genomics Global Neurodegenerative Disease Summit, we will bring the global neuroscience community together virtually to support one another, share research findings, and advance discovery to increase our understanding of neurodegenerative disease development, progression, and treatment.
To learn more about research from Dr. Ip and other leading neuroscientists, register today!
References:
- Lau SF, et al. Single-nucleus transcriptome analysis reveals dysregulation of angiogenic endothelial cells and neuroprotective glia in Alzheimer's disease. Proc Natl Acad Sci U S A 117(41): 25800–25809, 2020. doi: 10.1073/pnas.2008762117
- Reitz C, Brayne C & Mayeux R. Epidemiology of Alzheimer disease. Nat Rev Neurol 7: 137–152, 2011. doi: 10.1038/nrneurol.2011.2
- Chen Y, Fu AKY & Ip NY. Synaptic dysfunction in Alzheimer’s disease: Mechanisms and therapeutic strategies. Pharmacol Ther 195: 186–198, 2019. doi: 10.1016/j.pharmthera.2018.11.006
- Liddelow SA, et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541: 481–487, 2017. doi: 10.1038/nature21029
- Mathys H, et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570: 332–337, 2019. doi: 10.1038/s41586-019-1195-2
- Grubman A, et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat Neurosci 22: 2087–2097, 2019. doi: 10.1038/s41593-019-0539-4
- Wang X, et al. Deciphering cellular transcriptional alterations in Alzheimer’s disease brains. Mol Neurodegener 15: 38, 2020. doi: 10.1186/s13024-020-00392-6
