Inside (and outside) childhood cancers: resolving tumor heterogeneity and the tumor microenvironment for clues to improved therapies
September is National Childhood Cancer Awareness Month in the United States. To help raise awareness, we wanted to take this opportunity to dig into some of the latest research on pediatric cancers, including recent studies of pediatric ependymoma, a cancer that affects the brain and spinal cord, and B cell acute lymphoblastic leukemia (B-ALL). With the insights provided by single cell RNA-sequencing into tumor heterogeneity and the cellular composition of the tumor microenvironment, researchers are gaining more comprehensive knowledge of the ‘ins and outs’ of childhood cancer as they search for better therapies.

Though childhood cancer is rare, cancer is the most common cause of death by disease in children and approximately 300,000 children are diagnosed with cancer globally every year. Although survival rates have improved dramatically thanks to scientific and medical research, there’s still a lot we don’t know. And, unfortunately, almost all childhood cancer survivors will go on to suffer lifelong health complications due to cancer treatment, making the search for safer and more targeted therapeutic options of paramount importance.
Like most tumors, intratumoral heterogeneity and changes in the tumor immune microenvironment can impact disease trajectory and treatment response. Here, we’ll take a closer look at two recent studies that used single cell analysis to investigate the progression and therapeutic response of two types of childhood cancer. And we’ll explore the power of single cell resolution to give researchers a more complete perspective on childhood cancer—both inside and outside the tumor.
Cellular heterogeneity confounds traditional tumor subtyping
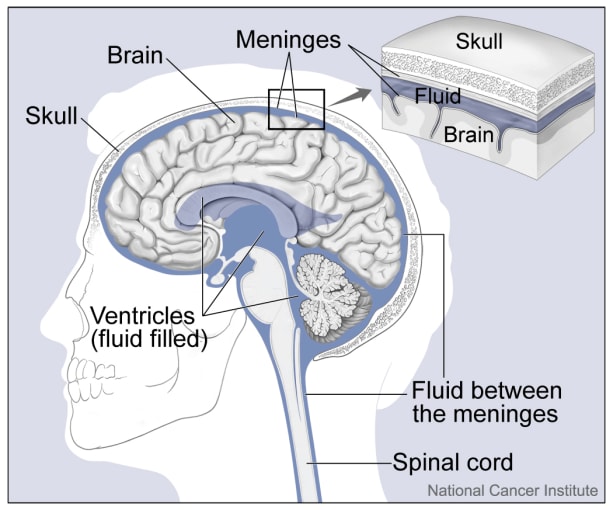
Ependymoma is a brain cancer more common in children than adults that affects the ependymal cells that line the ventricles and passageways of the brain and spinal cord. More than 50% of children with ependymoma will die from the disease, and relapse rates are high, with 70% of patients relapsing within 10 years. Four distinct subtypes of ependymoma have been identified based on bulk transcriptomic and methylomic studies. In particular, posterior fossa A (PFA) ependymomas are more likely to arise in younger children and have a poorer prognosis than ependymomas categorized as posterior fossa B (PFB). Given the high level of intratumoral heterogeneity observed in other brain tumors such as glioblastoma, a group of scientists from the University of Colorado Denver led by Andrew Donson set out to reveal the cellular composition of different ependymoma subtypes to see if bulk classification schemas could be improved (1).
The authors used Single Cell Gene Expression to analyze 26 childhood ependymoma samples. Using cluster analysis, they identified six major cellular subpopulations in tumors classified as PFA. Intratumoral heterogeneity was high. Each PFA sample included cells from at least two different cellular subpopulations, and half of the samples were made up of cells from all six subpopulations. A few of the subpopulations, including “undifferentiated ependymoma cells-1” (UEC-1), had never been described before.
To understand how cellular heterogeneity may be impacting tumor progression, Donson’s group used deconvolution of bulk transcriptomic data for 121 primary PFA ependymoma samples to estimate the proportion of each subpopulation within individual tumors. Although no change in overall survival is observed between PFA subtypes based on bulk methylome assignment, tumors with a higher-than-median fraction of UEC-1 had a significantly decreased survival rate compared to tumors with a lower fraction of UEC-1.
To understand how UEC-1 could promote cancer progression, the researchers returned to their single cell data. Using trajectory analysis, they ordered PFA cells in pseudotime and observed a divergent lineage trajectory, where UEC-1 could give rise to two distinct lineages, suggesting they may be the PFA cancer stem cell population. The authors suggest that two of the subpopulations they identified, including UEC-1, could be the drivers behind PFA relapse. They propose specifically targeting these cells to improve therapeutic outcomes and point out that “these subpopulations harbor distinct and clinically relevant biologies and, yet, were not apparent from bulk-tumor transcriptomic analyses, obscured by the overpowering transcriptomic profiles” of more prevalent subpopulations (1). In this way, a deeper dive inside pediatric ependymomas revealed previously unknown heterogeneity that may help researchers identify further treatment approaches.
Understanding the cellular context of cancer to identify new treatment options
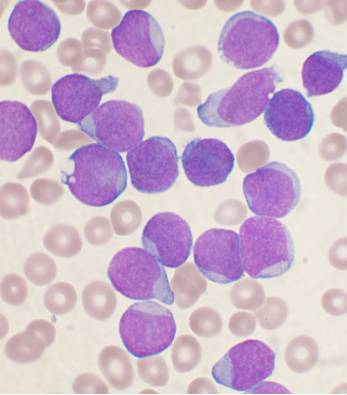
The most common childhood cancer is acute lymphoblastic leukemia, or ALL. In ALL, the bone marrow makes too many immature lymphocytes, resulting in susceptibility to infection, anemia, and easy bleeding. More than 10% of children with B cell ALL, in which excess numbers of immature B cells are produced, will suffer relapse and succumb to the disease. Although it is known that B-ALL transformation affects the bone marrow immune microenvironment, the manner in which this occurs and theextent of the resulting changes is not clear. To address this question, a group of researchers led by Iannis Aifantis at New York University School of Medicine used single cell RNA-sequencing and single cell identification of extracellular proteins (enabled here with CITE-seq, but now supported via Single Cell Gene Expression with Feature Barcode technology) to follow cellular changes in the bone marrow and peripheral blood of healthy patients and patients with B-ALL at diagnosis, remission, and relapse (2). To account for the predominant phenotype of B-ALL, namely a marked expansion in B cells, and enable discovery and analysis of non-leukemic cells, bone marrow samples were first flow sorted for B-cell and non–B-cell populations before being mixed at a 1:5 ratio.
Using single cell RNA-seq of 97,456 patient cells at diagnosis, remission, and relapse, the authors discovered that non-classical monocytes were enriched in leukemia samples at diagnosis and relapse. They then used CITE-seq to confirm the existence of leukemia-associated non-classical monocytes based on canonical protein markers. To understand whether the increase in non-classical monocytes had functional consequences, they looked at a cohort of 568 pediatric B-ALL patients with PBMC cell counts determined at diagnosis and throughout the course of therapy. Patients with abnormally high numbers of monocytes at diagnosis had inferior overall survival rates independent of other risk factors. This trend persisted in a study of adult B-ALL patients (2).
Armed with this knowledge, Aifantis and his collaborators next tested a potential therapeutic treatment for B-ALL. First, they confirmed an increase in non-classical monocytes after B-ALL transplantation in a mouse model. Using an antibody to block CSFR1, a method that has been shown to inhibit non-classical monocyte maturation, they treated leukemia-bearing mice with either a control antibody or anti-CSFR1 for 12 days followed by control treatment or chemotherapy. While anti-CSFR1 had minimal effect on disease outcome by itself, anti-CSFR1 in combination with chemotherapy enhanced overall survival and reduced the percentage of mice suffering relapse (2).
This study highlights the impressive strides that can be made when both clinical data and single cell analysis are available, enabling the investigation of novel treatment options that could someday enter the clinic and improve childrens’ lives. And it confirms the need to look outside the tumor itself, to the tumor context, immune–cancer cell interactions, and more, to gain a comprehensive understanding of the underlying biology driving childhood cancers.
We’re honored that our technology can support these incredible research projects, and the noble search for improved therapies. In honor of Childhood Cancer Awareness Month, we celebrate the many scientists, doctors, families, and children who are part of this larger story, and the larger fight against pediatric cancers.
References
- A Gillen et al., Single-Cell RNA Sequencing of Childhood Ependymoma reveals Neoplastic Cell Subpopulations That Impact Molecular Classification and Etiology. Cell Rep. 32, 108023 (2020).
- M Witkowski et al., Extensive Remodeling of the Immune Microenvironment in B Cell Acute Lymphoblastic Leukemia. Cancer Cell. 37, 867–882 (2020).
