What lies beneath: Uncovering mechanisms of selective vulnerability in Alzheimer’s disease
After facing significant setbacks in Alzheimer’s disease (AD) drug development (1), many scientists are going back to basics and aiming for a more fundamental understanding of disease mechanisms, keen to discover early changes at the level of individual cells.
While many researchers have chosen to focus on non-neuronal cell types and their contributions to degenerative disease (2), neuronal populations have received less attention. A fuller understanding of these cells could provide critical insights into this complex disease and shed light on new potential drug targets.
Single cell resolution: A transformative look at disease progression
Neurodegenerative diseases, including AD, are driven by selective vulnerability in neuronal populations, wherein a cell’s reaction to injury or disease varies based on its location and subtype (3). While previous work has examined this differential susceptibility and resilience at the surface level, data at the cellular level has been lacking. With the recent availability of technology that enables high-throughput single nucleus RNA-seq (snRNA-seq), Kampmann et al. at UCSF investigated the molecular profiles of neuronal subpopulations across disease progression (4).
Where earlier research focused merely on differences between healthy and diseased tissue, Kampmann’s team used single nucleus transcriptomics to closely examine progressive changes across different stages of the disease. Amyloid plaques and neurofibrillary inclusions are hallmarks of AD, the latter of which is measured by the Braak staging system, and correlates with clinical cognitive decline. The authors reasoned that profiling samples across different Braak stages would allow them to identify distinct molecular signatures that illuminate changes in neuronal subpopulations and cell type abundances that specifically characterize various time points in AD progression.
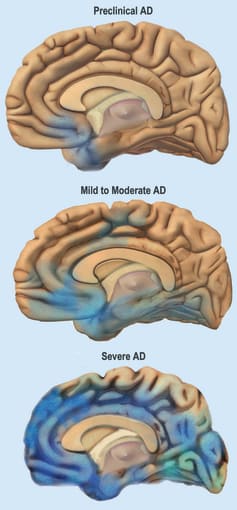
To accomplish this task, the researchers used Chromium Single Cell Gene Expression to perform snRNA-seq for transcriptomic profiling of postmortem brain tissue from ten male patients displaying a lack of AD-type tau neurofibrillary pathology (Braak stage 0) or early- and late-stage pathology (Braak stages 2 and 6). Nuclei were extracted from matched samples of the entorhinal cortex (EC) and superior frontal gyrus (SFG), regions of the brain where neurofibrillary inclusions and neuronal loss occur early and late in AD, respectively.
Clusters were assigned to broad cell types based on known markers, then proportions of identified cell types were assigned to each patient and assessed to determine how they changed across Braak stages and, therefore, AD progression. A drop in the relative abundance of excitatory neurons was seen in the EC at Braak stage 2 and in the SFG at Braak stage 6. These findings aligned with previous observations indicating earlier susceptibility of the EC versus the SFG and higher vulnerability of excitatory versus inhibitory neurons.
Identifying neuronal subpopulations and their fates in AD pathology
Subclustering of excitatory neurons in the entorhinal cortex (EC) and superior frontal gyrus (SFG) revealed nine subpopulations, each exhibiting distinct expression patterns of genes specific to the EC. Three of these subpopulations showed compelling evidence of early depletion, with a 50–60% drop in relative abundance in Braak stage 2 compared to stage 0. These three groups of cells expressed genes associated with EC layer 11, a site of preferential accumulation of tau neurofibrillary inclusions in early-stage AD. However, this gene expression signature was also seen in subpopulations that were not depleted across later Braak stages.
The search for molecular markers specific to vulnerable excitatory neurons then shifted to differential expression patterns between pairs of subpopulations. A closer look at these transcript levels revealed a select set of genes expressed by no more than four subpopulations. Among them, RAR-related Orphan Receptor B (RORB) was specifically expressed by two of the previously identified subpopulations, as were two noncoding transcripts. These vulnerable neurons also displayed lower expression of genes involved in multiple signaling pathways.
A comparison of gene expression across Braak stages 0 and 6 for all EC excitatory neuronal subpopulations unsurprisingly showed the widest disparity in differentially expressed genes and supported the recent findings of Marinaro et al., who observed the downregulation of synapse-related genes in a snRNA-seq study of familial monogenic AD (5).
Confirming selective vulnerability markers across brain regions
An examination of excitatory neurons in the superior frontal gyrus (SFG) revealed that two subpopulations were only depleted in late-stage AD and shared the same three vulnerability markers identified in the entorhinal cortex (EC). The decrease in relative abundance at a later point in disease progression is supported by the observed timing of inclusions in this brain region. Taken together, the authors suggest that these data point to similar mechanisms underlying selective vulnerability across different regions of the brain and a possible focus for future research.
Building a foundation for clinical success
The complex nature and progression of AD, paired with the low success of drugs intended to target plaque and tangle accumulation (1), means that expanding the breadth and depth of our basic understanding of the brain is critical in the search for an effective treatment. From an enhanced knowledge base, scientists can acquire a more comprehensive understanding of the mechanisms involved in selective neuronal changes and, therefore, greatly expand the list of potential drug targets. While a solution is currently elusive, gaining a firmer grasp on the functional ramifications of transcriptomic changes may very well lead to therapeutic success.
References
- https://www.advisory.com/daily-briefing/2020/02/12/alzheimers-disease
- Olah M et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat Comm 11: 6129, 2020.
- Mattsson N et al. Selective vulnerability in neurodegeneration: insights from clinical variants of Alzheimer’s disease. J Neurol Neurosurg Psychiatry 87: 1000–1004, 2016.
- Leng K et al. Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease. Nat Neuro 24: 276–287, 2021.
- Marinaro F et al. Molecular and cellular pathology of monogenic Alzheimer’s disease at single cell resolution. bioRxiv, 2020. DOI: 10.1101/2020.07.14.202317
