Visium CytAssist Spatial Gene Expression Library, Human Lymph Node (FFPE)
Spatial Gene Expression dataset analyzed using Space Ranger 3.1.3
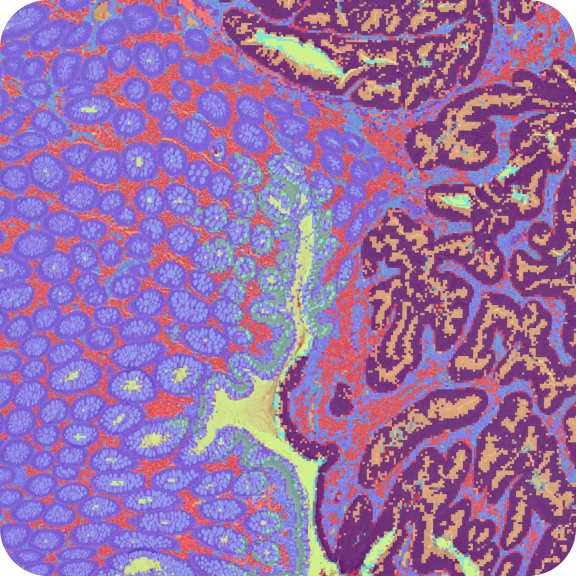
Learn about Visium Analysis
Biomaterials
FFPE human lymph node tissue was obtained from Avaden Biosciences.
Sample preparation
A 5 µm section was taken with a microtome (Epredia HM355S). Tissue block preparation, sectioning, H&E staining, and imaging followed the Visium CytAssist Spatial Gene Expression for FFPE – Tissue Preparation Guide (CG000518).
Imaging
- Image type: H&E
- Microscope: Olympus VS200 Slide Scanner
- Objective magnification: 10X
- Numerical Aperture: 0.8
- ScopeLED light source: VS200 LED, Olympus integrated bright field source
- Camera: iDS VS-264C, Olympus scanner integrated camera
- Exposure: 500 microseconds
Assay workflow
Probe hybridization, probe ligation, slide preparation, probe release, extension, and library construction followed the Visium CytAssist Spatial Gene Expression Reagent Kits User Guide (CG000495).
- Slide serial number: V43J30-328
- Area: A-1
- Instrument: Visium CytAssist
- Probe set: Visium Human Transcriptome Probe Set v2.0
Sequencing
- Indexing: Dual index plate TS set A; sample index C12
- Sequencing instrument: Illumina NovaSeq 6000
- Sequencing configuration: 28 bp read 1, 90 bp read 2, 10 bp i7 sample index, and 10 bp i5 sample index
- Sequencing depth: 275.8 million read pairs
Analysis
Space Ranger v3.1.3 was used to map reads to the reference, detect tissue, align the data to the microscope and CytAssist images, and output feature-barcode matrices for further analysis.
How to view data
To get started, download Loupe Browser v8.1 to explore the Loupe file, or read more about the other Space Ranger outputs.
This dataset is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) license. 10x citation guidelines available here.
