There are several ways to use the 10x Genomics Cloud Analysis platform for your data analysis: 1) web browser app, 2) command line (CLI) interface, and 3) prompt-based interface.
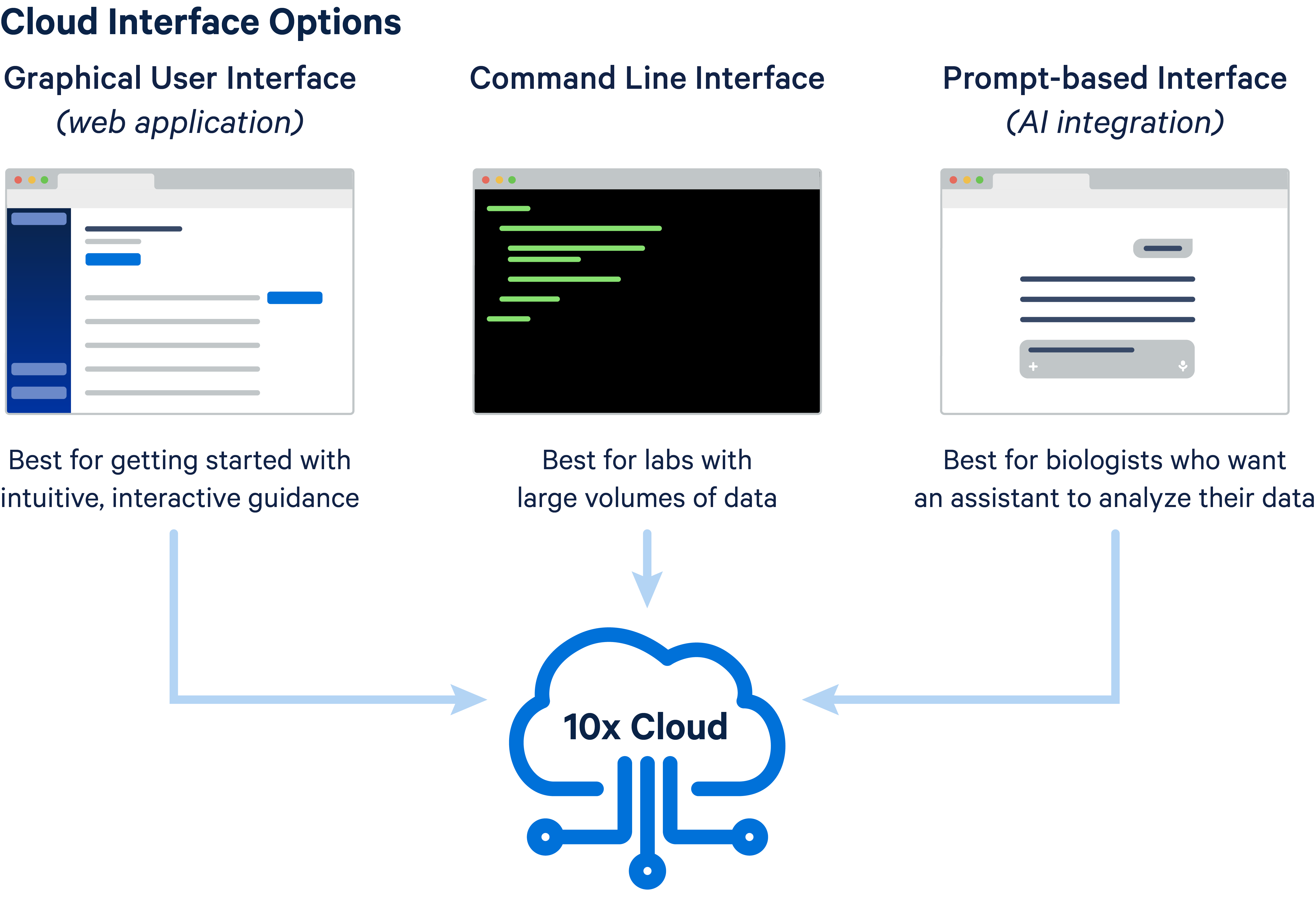
All methods begin with 10x Cloud account creation. After your account is created, the general steps include uploading inputs, creating your analysis, and downloading outputs. The way these steps are run and the options available depend on the method you select.
To help you decide the best method for your use case, review these considerations:
| Cloud method considerations | Web browser app | Command Line (CLI) interface | Prompt-based interface |
|---|---|---|---|
| Batch analysis set up | Manual set up only | Yes, via scripting | Yes, via prompts |
| Interface type: graphical user interface (GUI) or command-line (CLI) | GUI | CLI | Both GUI and CLI |
| Normalization and/or batch correction workflow (SCTransform and Harmony) support | Yes (Chromium only) | Not available | Not available |
- Refer to the supported products page for a list of supported data and library types, as well as pipeline version details.
- Refer to these tables for supported single cell sub-pipelines and supported spatial sub-pipeline.
Once you have decided on a method, proceed to the corresponding documentation for usage details: