A basic introduction to linked-reads
Standard short read sequencing provides accurate base level sequence to provide short range information, but struggles to provide long range information. This means that standard sequencing and analysis approaches typically do well at calling single nucleotide variants (SNVs) but fail to robustly identify the full spectrum of structural variation seen in an individual genome. A novel data type known as Linked-Reads utilizes molecular barcodes to tag reads that come from the same long DNA fragment.
Linked-Reads provide the long range information missing from standard approaches. By adding a unique barcode to every short read generated from an individual molecule, you can link the short reads together.
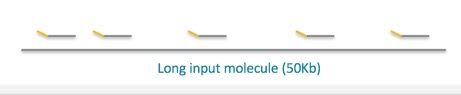

Note in the figure above, several reads (grey lines) are generated from this long input molecule and they each contain the same barcode (gold line). This allows you to deduce that these reads came from the same molecule.
Many people wonder why we don’t fully saturate the molecule with barcoded reads, to make a synthetic long read. The synthetic long read approach increases sequencing cost and typically means that you have overall less physical coverage for equivalent sequence coverage and cost.
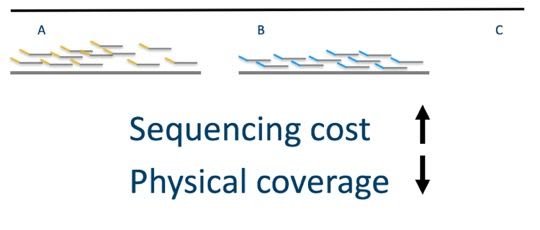

In the image above, we see two long molecules with lots of read coverage, but we still lack the ability to link the three loci (A, B and C).
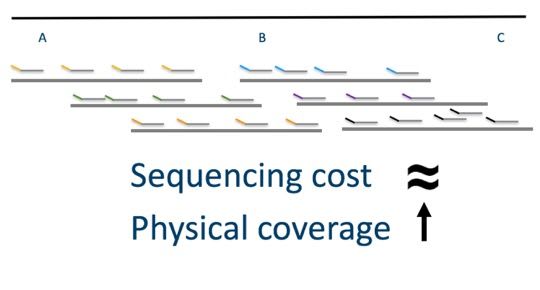
By sequencing less deeply, we can incorporate more long molecules into the system for equivalent sequence depth. This gives us the power we need to link distant loci. When following the standard Chromium Genome and sequencing to 30x coverage you have, on average, 150x physical coverage. This degree of long range information enables reconstruction of long range haplotypes, power to call complex structural variants and enables the ability to perform de novo genome assembly.
Learn more about all our Linked-Read resources, including seminars and application notes here:
