Human Skin Data with Xenium Human Multi-Tissue and Cancer Panel
In Situ Gene Expression dataset analyzed using Xenium Onboard Analysis 1.9.0

Learn about Xenium analysis
Overview
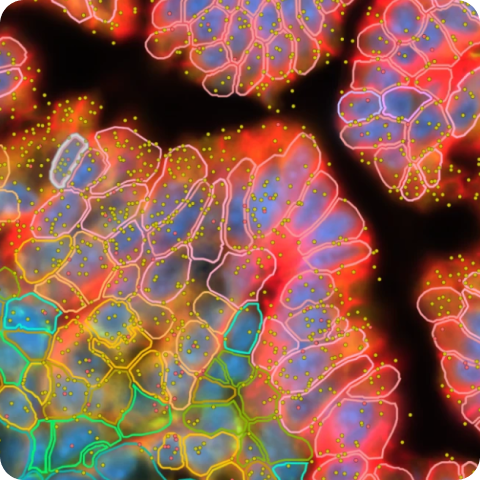
Xenium In Situ Gene Expression data for adult human skin sections using the Xenium Human Multi-Tissue and Cancer Panel.
How to view data
Interactively explore data with Xenium Explorer by downloading the Xenium Output Bundle (or Xenium Explorer subset) file. The subset bundle contains the experiment.xenium, morphology_mip.ome.tif, analysis_summary.html, cells.zarr.zip, cell_feature_matrix.zarr.zip, transcripts.zarr.zip, and analysis.zarr.zip files. See the Getting Started with Xenium Explorer page for more details. Follow these instructions to view the post-Xenium H&E image or image alignment file in Xenium Explorer.
Biomaterials
FFPE-preserved tissues were purchased from Avaden Biosciences (non-diseased skin).
Tissue preparation
Tissues were prepared following the Xenium In Situ for FFPE - Tissue Preparation Handbook (CG000578). Additional guidance for sectioning skin samples is provided here.
Post-instrument processing followed the Demonstrated Protocol Xenium In Situ Gene Expression - Post-Xenium Analyzer H&E Staining (CG000613).
Gene panels
The Xenium Human Multi-Tissue and Cancer Panel (377 genes) was pre-designed by 10x Genomics. The panel design was informed using single cell RNA sequencing data curated and reprocessed for standardization by the Human Protein Atlas. Genes were chosen to accurately type cells, and identify select immune, proliferation, and tumor markers, in human breast, lung, skin, liver, colon, kidney, lung cancer, and heart.
Xenium Analyzer
The instrument run was performed following the Xenium Analyzer User Guide (CG000584). The on-instrument analysis was run with Xenium Onboard Analysis version 1.9.0. The analysis summary for both samples shows the "Low decoded transcripts per 100 µm2" warning. This is expected because the epidermis contains very high transcript counts for genes on the panel, but the other cell types in skin have lower counts by comparison. In these two samples, the epidermis is only a fraction of the total surface area of the samples, thus contributing to an overall low number of decoded transcripts per 100 µm2.
| Metric | Sample 1 | Sample 2 |
|---|---|---|
| Median transcripts per cell | 25 | 17 |
| Cells detected | 68,476 | 90,106 |
| Decoded transcripts per 100 µm² | 9.8 | 8.7 |
| Total high quality decoded transcripts | 2,113,721 | 1,797,665 |
| Region area (µm²) | 72,922,736 | 43,243,357 |
This dataset is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) license. 10x citation guidelines available here.