Forging new connections at Society for Neuroscience 2023
As a neuroscientist, there’s nothing like attending the Society for Neuroscience conference every year. This event brings together 30,000 neuroscientists for the chance to meet and discuss everything from neuroscience education to ethics. It allows us to showcase our most up-to-date tools, whether it’s the latest in single cell spatial imaging, advances in spatial transcriptomics, or single cell RNA-seq from fixed tissue. Most of all, it allows us to connect with the neuroscientists who are using these tools to drive research into new and exciting frontiers.
One such neuroscientist is the subject of this blog: Charles Wang, MD, PhD, MPH, at Loma Linda University’s School of Medicine. He won the 10x Genomics poster contest with his entry, “Single-nucleus multiome and spatial transcriptomics reveal abnormal neural development in rat brain after prenatal e-cigarette exposure.” We were thrilled to be able to sit down for a Q&A with him at the conference and discuss his past, present, and future body of work.
“What I see is, with all these single cell sequencing and spatial genomics, maybe even single cell proteomics, the possibility that—if we can do everything with a single cell—we will truly understand biology, especially with spatial genomics.” –Dr. Charles Wang
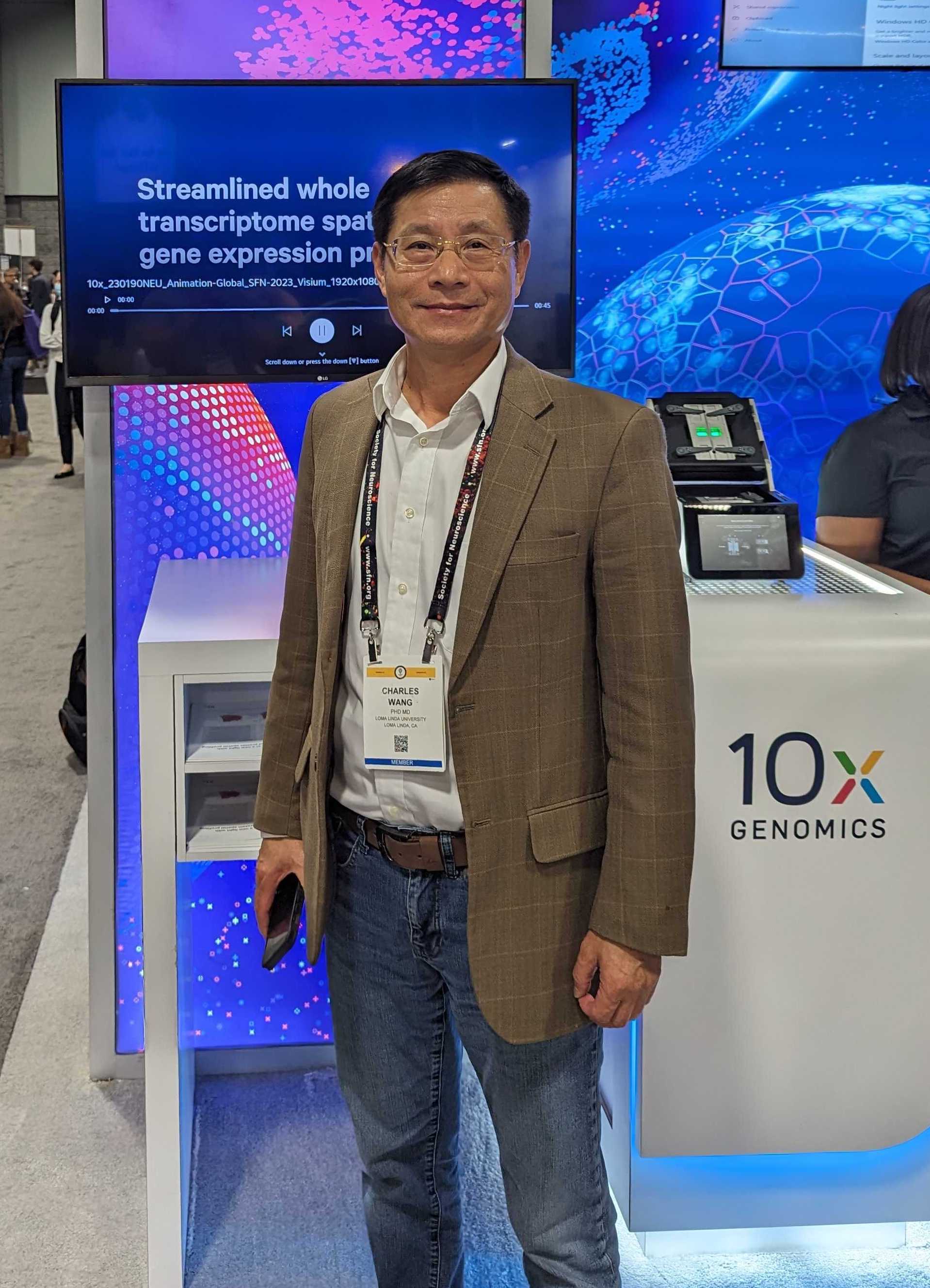

Can you give us an overview of what led you down the path to this kind of research?
I'm a genomic person, I'd say. My expertise is really in the genomic, epigenomic, and single cell sequencing methods. I've been running the genomics core or center lab for more than 20 years and I used to work at the Cedars Sinai Medical Center as the transcriptional genomic core director with a joint appointment as an assistant/associate professor at UCLA School of Medicine.
10 years ago, I moved to Loma Linda University to set up a genomics center from scratch, and, even at that time, I was extremely interested in single cell sequencing. However, at that time, single cell sequencing didn't work so well; so I was so glad to get the 10x single cell sequencing technology in 2016.
I was also one of the key leaders and played a leadership role for the FDA's sequencing quality control (SEQC) projects. So back in 2017, I thought about benchmarking single cell sequencing across multiple centers and multiple platforms. Then I challenged myself and designed and carried out a multicenter benchmark study on single cell sequencing, using the two reference cell lines that we used for benchmarking the whole genome sequencing technologies.
I had three papers published in Nature Biotechnology in 2021*, but the single cell paper was the very first one published online (in 2020) which included several single cell technologies. The single cell sequencing multicenter study paper won the best paper award at the 2021 MAQC society annual meeting. I was even invited to give a talk by NCI in their cancer data seminar.
**Author’s note: The papers mentioned above can be found here, here, and here.*
It's nice seeing that kind of rigor applied to any new technology. New tools let us do new things, but it's always a question of how you can really think of new technologies, how can you really trust them, so thank you for doing those types of studies.
That's what we do. We want to benchmark and see how reliable, and how consistent, tools are across different centers, different labs, etc. So, for that paper, we not only benchmarked the technologies, but we actually saw how bioinformatics analysis is vital. It's critical. So there's a very nice story there in terms of how to handle single cell data provided from different technologies, different labs, etc.
I remember trying to learn bioinformatics from scratch. It was...challenging to say the least. So it's nice having a lot more of these custom built tools available that have been validated by outside parties that we can actually trust! How did that lead to the research you did for the poster and your current research projects?
Parallel to the multicenter single cell RNA sequencing benchmarking study, I had another project using a prenatal electronic cigarette (e-cig) exposure animal model. So I took your single cell sequencing technology, your Multiome that gives both single nucleus RNA-seq and ATAC-seq, and used it to analyze the rat brains prenatally exposed to e-cig. That study generated some major hypotheses that led to us applying your spatial technology and helped generate data for our grant.
Is there anything you can say about those findings and what drove you to use spatial technologies?
When we used the 10x Genomics Multiome Single Cell sequencing technology, I really loved it. We took the prenatal e-cigarette-exposed fetal brains at postnatal day 7, took specific cortical and hippocampal regions, and then isolated nuclei. Then we did single nucleus RNA-seq and single nucleus ATAC-seq. The major discovery was we noticed that the ratio between two types of neurons, excitatory and inhibitory neurons, was totally disrupted.
This led us to the hypothesis that prenatal e-cigarette exposure induced the excitatory-inhibitory imbalance through epigenetic mechanisms that somehow affect the neural progenitor cells. We really want to get a comprehensive genomic understanding of abnormal brain development with prenatal e-cigarette exposure.
We’re going to use single cell sequencing and spatial genomics—I proposed a lot of Visium experiments—across several time points. And we want to use spatial genomics to see which specific cell type, which specific location, might be more sensitive during early embryonic or postnatal brain development. Of course, we’ll also simultaneously do single cell sequencing. This approach would really allow us to understand the epigenetic reprogramming in this animal model.
How have you seen the single cell landscape change over the course of your career?
I’ve really witnessed the advances in cutting-edge technologies from the human genome sequencing project, to genome microarrays, to next-generation sequencing, single cell sequencing, spatial genomics. I’m so excited. I’m a technology-driven person, and I don’t invent these methods but I apply them to my projects very fast.
What I see is, with all these single cell sequencing and spatial genomics, maybe even single cell proteomics, the possibility that—if we can do everything with a single cell—we will truly understand biology, especially with spatial genomics. So I think all these new tools really empower us to push life sciences to a totally different level. Just like the Human Genome Project 20 years ago, more than $10 billion dollars and 10 years, and now it’s so much easier.
That’s one of my favorite examples of progress. People forget: it’s been only 60 years since we discovered the structure of DNA, 35 years since PCR, 15 years since next-gen sequencing, 10 years for single cell sequencing. I can’t wait to see what researchers like you will do in the next 10 years. And speaking of: is there anything you’d really like us and the readers to know about your work, or anything you’ve done that you’d like us to amplify?
Really, the multicenter studies since they heavily involve 10x single cell sequencing platforms, and to promote our paper and promote the study. I will say that single cell sequencing technology is so powerful, it’s very powerful, and we’ve done a very nice comprehensive benchmarking study that can be found in Nature Biotechnology. Using the single cell sequencing technology in a prenatal animal model let us generate some very exciting data that led to a new hypothesis, then led to a major genomic and epigenomic U01 [NIH] grant.
I also do feel very excited, since I happen to know that you guys will release high-definition Visium. Visium has worked so well for us, and that’s one thing I’m really excited for; and the other is my (Visium) CytAssist that I got a couple years ago. It’s a nice design—easy and user friendly.
Thank you so much for your time, and congratulations again on winning the poster contest!
Shining a spotlight on the runners-up: Ischemic injury, stroke, sleep deprivation, and psychiatric disorders
We’d also like to congratulate and highlight the work from our four runners-up, Rachel Kim (New York University School of Medicine), Pares Habib, PhD (Stanford University), Yann Vanrobaeys (University of Iowa), and Ahyeon Hwang (University of California, Irvine). We’ve provided brief synopses of their abstracts, and encourage you to check out their work!
Kim RD, et al. “Molecular heterogeneity of ischemic injury-induced reactive astrocytes.” Kim et al. used Visium spatial transcriptomics to characterize astrocytes following ischemia and found temporal, cellular, and spatially restricted astrocytic subtypes that could play functional roles in injury stabilization and recovery. Read the abstract.
Habib P, et al. “Spatial mapping of stem cell-induced transcriptional alterations in stroke-injured rat brains.” Building on a prior clinical trial examining stem cell treatments in human stroke patients, Habib et al. used Visium spatial transcriptomics to characterize specific and spatial gene signatures associated with stroke injury, stem cell grafts, and other phenomena. Read the abstract.
Vanrobaeys Y, et al. “Spatial transcriptomics reveals unique gene expression changes in different brain regions after sleep deprivation.” In their study, Vanrobaeys et al. leveraged spatial transcriptomics in order to see the impact that sleep deprivation has across the entire brain, rather than focusing on individual regions that may not tell the whole story. Using this approach, they defined multiple regions with the greatest transcriptional impacts—specifically, hippocampus, neocortex, thalamus, and hypothalamus—and were able to get a more holistic view of the brain in sleep deprivation. Read the abstract.
Hwang A, et al. “Single cell transcriptomic and epigenomic dynamics of >1M nuclei from the human PTSD brain.” Hwang et al. examined both the transcriptomes and epigenomes at single cell resolution from patients with post-traumatic stress disorder (PTSD), patients with major depressive disorder (MDD), and healthy controls. Using this approach, they were able to better characterize the regulatory network and cellular basis of these complex disorders in the dorsolateral prefrontal cortex (DLPFC), a brain region central to the regulation of emotion and cognition. Read the abstract.
We had a great time seeing everyone at SfN 2023 and can’t wait to see you next year in Chicago. In the meantime, stay up to date with the latest in spatial transcriptomics with Visium + protein, explore Chromium single cell multiomics, or find out how Xenium single cell spatial is unlocking new avenues of inquiry. Until next time!
