Global Immunology Virtual Symposium: Meet three of the researchers making a mark on immunology
If we had to pinpoint one lesson to take from the past year, it might be something like this: immunological research is essential. That’s why we recently held the Global Immunology Virtual Symposium, an event gathering some of the leading minds in immunology as they shared their latest discoveries, data, and innovative methods.
We were blown away by all of our presenters. Their fascinating research represents the future of immunology, and we’re more convinced than ever that the future is looking bright. We’ve highlighted three of the presentations below, so keep reading to get a peek. Then, watch all the presentations on-demand here.
Integrating single cell transcriptomics with alloreactive TCR sequencing in human transplant recipients
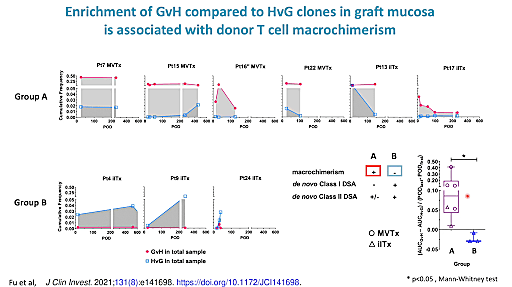
Our plenary speaker Megan Sykes, MD, Director of the Columbia Center for Translational Immunology at Columbia University, started things off, discussing her research into transplantation dynamics and allograft tolerance induction. Currently, the transplantation field is limited by drug treatment-related complications and chronic rejection. However, both of these limitations can be avoided by induction of immune tolerance. In other words, the goal is long-term graft acceptance without the need for immunosuppressive therapy and with an otherwise competent immune system.
Dr. Sykes’ studies focus on hematopoietic cell transplantation (HCT), which, in an earlier study, she found to be a powerful method of inducing tolerance in animal models. Unfortunately, translating this to human patients is far from simple as treatments must be minimally toxic, while still being strong enough to induce host and donor cells to mix across HLA barriers (mixed chimerism). Additionally, graft-vs-host disease (GVHD), a very common complication in transplantation, is a serious problem, even in patients who simply need an organ allograft and do not otherwise have malignant disease. Following murine trials in which Dr. Sykes and her colleagues found that GVH reactions (GVHR) were not necessarily equivalent to GVHD and could create graft vs leukemia or lymphoma (GVL) effects without GVHD in certain cases, Dr. Sykes and her colleagues conducted a series of studies and clinical trials investigating mixed chimerism and its role in allograft tolerance. Using Chromium Single Cell Gene Expression and Immune Profiling, Dr. Sykes was able to dissect the mechanisms underlying multi-lineage chimerism and successful hematopoietic stem and progenitor cell engraftment in human intestinal transplantation. Importantly, her discoveries suggest a possible novel approach to achieving intestinal allograft tolerance.
Watch Dr. Sykes’ presentation here →
Systems immunology approach reveals novel mechanisms of lung pathology in severe COVID-19
Before the COVID-19 pandemic, Eliver Ghosn, PhD, Associate Professor at Emory University, and his colleagues were focused on immune system development and function, studying tissue-resident immune cells that form early during fetal life and persist throughout adulthood in both humans and mice. Looking at the human lung, they found two distinct lineages of immune cells—tissue-resident macrophages originating from embryonic life and infiltrating monocytes from adult bone marrow stem cells. Their primary research question centered on these different lineages and their response to viral infection.
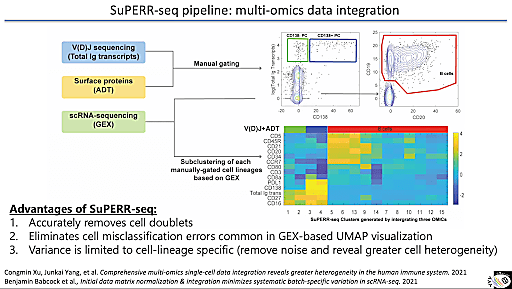
After the emergence of COVID-19, they still sought to answer their main research question, however, they pivoted slightly, shifting their focus to the lung immune system response to COVID-19. Dr. Ghosn and his team used a multimodal pipeline they call SUPERR-seq, combining V(D)J sequencing and protein analysis with SARS-CoV-2-inclusive single cell RNA-sequencing (scRNA-seq) powered by Chromium Single Cell Gene Expression to detect immune cells affected by the virus in the lungs of severe COVID-19 patients. They uncovered a neutrophil-activated feedback loop in the lung that results in the exacerbated inflammatory response which characterizes severe COVID-19. Maintaining their systems immunology focus, the team then set out to understand the origins of these neutrophils and the mechanisms by which they enter the lung, anticipating the possibility of targeting neutrophils before they enter the lung or preventing their migration altogether.
Learn more about this study →
Development of multimodal single-cell sequencing for rhesus macaque
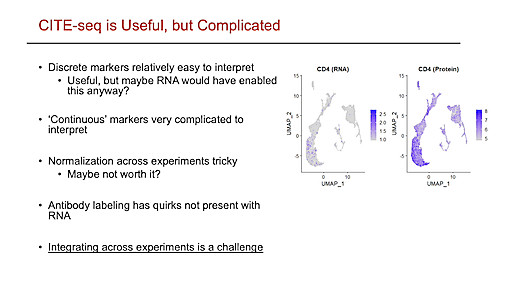
Benjamin Bimber, PhD, Research Assistant Professor at the Oregon National Primate Research Center, is also applying a multimodal approach in his study of the immune system in rhesus macaques. Unlike human or mouse, macaque is not a particularly widely studied organism in this field. As a result, Dr. Bimber and his team are often working with precious, low-input samples, which is what first drew them to Chromium Single Cell Gene Expression. The Chromium platform has a high capture rate of up to 65% and provides robust data, even from limited samples. Another benefit of the Chromium technology is its multiomic capabilities, allowing Dr. Bimber and his colleagues to add on T-cell receptor sequencing and cell surface protein measurement to their single cell analysis. As a result, they can better understand immune dynamics in macaques, generating high-resolution multiomic data across tissues and over time. Now, having generated a large amount of single cell data across different tissues and individuals, they are working to build a macaque cell atlas, annotating molecular profiles gleaned from scRNA-seq data to their canonical cell immune phenotypes. This atlas will build a better foundation from which to analyze and compare data across studies, providing a more stable reference for researchers working with macaque samples. Dr. Bimber also expressed a hope that his team’s methods would help other researchers working outside of human and mouse models as they develop methods optimized for their own organisms of interest.
Watch Dr. Bimber’s talk here →
And visit our** Immunology hub** to learn more about how 10x Genomics technologies let you address the intricacies of the immune system.
