Building a complete picture of cancer: A team effort
We invite all cancer researchers to come together virtually as a global community to support each other, share research findings, and advance discovery. Explore the latest research with leading scientists at the Virtual Cancer Symposium, and find guidance for single cell sample preparation workflows that can empower you to build on their findings and resolve the complexities of cancer.
Whether deciphering the cellular features of the tumor microenvironment, or investigating the basis of therapeutic response, cancer researchers face the challenge of developing comprehensive knowledge of a complex and heterogeneous disease. Through collaboration and shared technological innovations, however, a more complete picture of cancer biology can emerge.
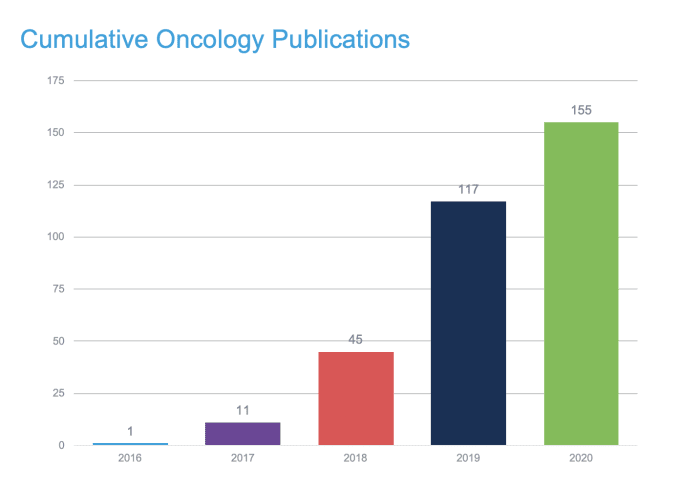
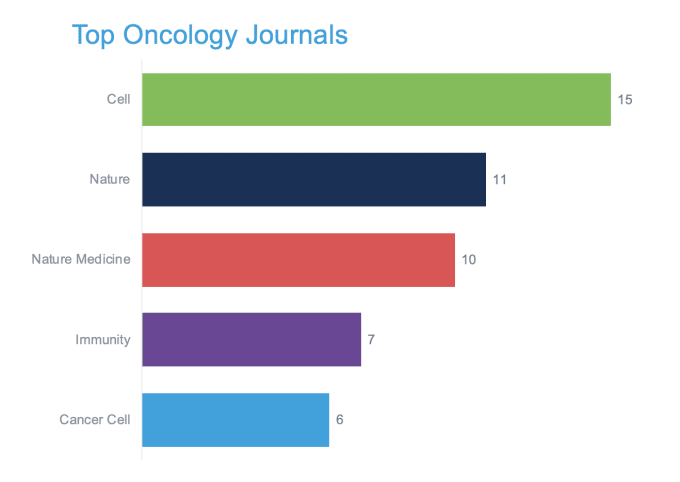
In that spirit, 10x Genomics invites you to join researchers from around the world at the Virtual Cancer Symposium, a week-long series of scientific presentations from leading researchers who have used single cell and spatial gene expression technology to advance our knowledge of cancer. Preview some of their research projects in the publications highlighted below, and find inspiration for your own studies.
Epigenomics and immune repertoire profiling unveil tumor-resident T-cell biology
Dr. Ansuman Satpathy, MD, PhD, Assistant Professor at Stanford University, has a long history with T cells. He and his colleagues at Stanford University, the Parker Institute for Cancer Immunotherapy, and other partner institutions have collaborated on 4 recent publications that focus on tumor-resident T-cell exhaustion and how immune checkpoint blockade therapies work to reinvigorate these cells. Leveraging single cell ATAC-seq and single cell immune profiling technology, Satpathy and his collaborators profiled T cells from various tissues, including human blood, site-matched tumors, and serial tumor biopsies in patients with basal and squamous cell carcinoma, before and after anti-PD-1 therapy. With access to the chromatin regulators of therapy-responsive T-cell subsets, and the ability to perform T-cell clone tracing, they were able to make some unexpected findings. Significantly, exhausted T cells were not reinvigorated by PD-1 blockade. Rather, tumor-resident T cells were epigenetically locked in an exhausted state, and the T cells mounting the anti-tumor response were actually new T cells expanded in the periphery.
Read more about these findings:
- A Satpathy et al., Massively Parallel Single-Cell Chromatin Landscapes of Human Immune Cell Development and Intratumoral T Cell Exhaustion. Nat. Biotechnol. (2019).
- K Yost et al., Clonal Replacement of Tumor-Specific T Cells Following PD-1 Blockade. Nat. Med. (2019).
T-cell exhaustion poses a major challenge to developing effective and sustainable cancer immunotherapies. With this challenge in mind, Satpathy and his colleagues have also explored the efficacy of various modes of T-cell engineering to alleviate exhaustion. In the first in-human, phase 1 clinical trial of its kind, one team of researchers evaluated the utility of CRISPR-edited T-cell therapy in refractory cancers. They knocked down three genes encoding T-cell receptors, including PDCD1, following the school of thought that PD-1 blockade can reinvigorate exhausted T cells. Leveraging data from 5' single cell RNA-sequencing (scRNA-seq) libraries, they were able to monitor T-cell stability and abundance in circulation over time. Building upon the findings of previous studies, they discovered that PD-1 deficient engineered cells were less abundant in circulation over time, indicating these cells were less capable of establishing memory. These findings will continue to guide future studies that aim to discover the right targets for engineering T cells.
Explore additional publications:
- R Lynn et al., c-Jun Overexpression in CAR T Cells Induces Exhaustion Resistance. Nature. (2019).
- EA Stadtmauer et al., CRISPR-Engineered T Cells in Patients with Refractory Cancer. Science. (2020).
Learn more about these crucial developments for cancer immunotherapy in Dr. Satpathy’s presentation at the Virtual Cancer Symposium. Register here →
How peripheral T-cell expansion affects clinical outcome
Recent findings about the interplay of immune checkpoint therapy and intratumoral T-cell exhaustion have clinical implications that are now being assessed by research teams at major pharmaceutical companies and in an expanded number of cancer types. Dr. Thomas Wu, MD, PhD, Principal Scientist at Genentech, led a study of T-cell clonal expansion in lung, colon, and renal tumors resected from patients following PD-1 therapy, and in normal adjacent tissue and peripheral blood from the same patients.
Using scRNA-seq and T-cell receptor (TCR) sequencing, they found expanded T-cell clonotypes in the tumors, normal adjacent tissue, and in peripheral blood samples. Critically, patients with expanded peripheral T-cell populations responded best to PD-1 blockade therapy, suggesting that, mechanistically, immune checkpoint blockade drives recruitment of novel, non-exhausted clones from sites outside the tumor rather than reinvigorating exhausted intratumoral clones. These findings not only build on work conducted by Dr. Satpathy and his colleagues, but also point to the possible diagnostic value of peripheral clonotypic expansion as a predictor of clinical response to immune checkpoint blockade.
Explore this study in more detail:
- T Wu et al., Peripheral T cell expansion predicts tumour infiltration and clinical response. Nature. (2020).
And register for Dr. Wu’s presentation at the Virtual Cancer Symposium →
Uncovering the role of intratumoral heterogeneity in therapy resistance
Beyond T-cell exhaustion, some innate characteristics of the tumor can influence therapeutic response. The trouble is, how do you find those specific molecular features in a sea of heterogeneous cancer cells?
Dr. Allison Stewart, PhD, Staff Scientist, and Dr. Carl Gay, MD, PhD, Assistant Professor, both of the Department of Thoracic/Head and Neck Medical Oncology at the University of Texas MD Anderson Cancer Center, sought to understand this problem in their study of small cell lung cancer (SCLC). SCLC is notorious for initially being highly responsive to chemotherapy treatment, and then overwhelmingly and rapidly resistant post-recurrence of disease. Leveraging scRNA-seq to study circulating tumor cell-derived murine xenografts, including models derived from both chemoresponsive and chemoresistant patient tumors, the research team uncovered coexisting subpopulations of cells with heterogeneous gene expression of therapeutic targets and potential resistance pathways. This finding suggests that multiple resistance mechanisms develop concurrently, and may even be attributed to selective pressure induced by monotherapy. If this is the case, a diverse, combination therapy strategy from the start of treatment could maximize efficacy before the emergence of this observed transcriptional heterogeneity.
Dr. Stewart and Dr. Gay will explore the clinical implications of their single cell data in a presenation at the Virtual Cancer Symposium. Register here, and explore their publication:
- CA Stewart et al., Single-Cell Analyses Reveal Increased Intratumoral Heterogeneity After the Onset of Therapy Resistance in Small-Cell Lung Cancer. Nat. Cancer. (2020).
Deciphering chemoresistance in breast cancer
Other cancer studies have benefited from the ability to uncover cell-type specific transcriptional programs underlying chemoresistance. Using patient-derived murine xenograft models of triple negative breast cancer (TNBC) and leveraging bulk and single cell RNA-sequencing, Dr. Alexander Swarbrick, PhD, Associate Professor at UNSW Sydney, and his research team determined that a Hedgehog ligand secreted byneoplastic cells reprograms cancer-associated fibroblasts (CAFs) to provide a supportive niche for the acquisition of a chemo-resistant, cancer stem cell phenotype. Conversely, they observed that treatment of CAFs with smoothened inhibitors (SMOi) can reverseCAF gene expression changes induced by Hh signaling. Their findings point to the power of single cell sequencing approaches to reveal crucial interactions between stromal and cancer cells in the tumor microenvironment, and characterize the cellular and molecular features that drive chemoresistance and present novel therapeutic opportunities.
Explore this study in more detail:
- A Cazet et al., Targeting stromal remodeling and cancer stem cell plasticity overcomes chemoresistance in triple negative breast cancer. Nat. Comm. (2018).
For his presentation at the Virtual Cancer Symposium, Dr. Swarbrick will continue this exploration of breast cancer, discussing how his research team integrated single cell and spatial gene expression data to gather new insights into the disease. Register here →
Our understanding of cancer is evolving and deepening as each new discovery builds upon its predecessors. As we anticipate the future of cancer research, collaboration and shared success stories, whether pertaining to best practices for sample preparation and tissue dissociation, or to innovative experimental applications of single cell and spatial gene expression technology, can only accelerate further discovery. So come to the Virtual Cancer Symposium ready to collaborate and find the support you need for your own study of cancer alongside a global cohort of scientific peers and 10x Genomics scientists.
