How to streamline sample prep with the Chromium Nuclei Isolation Kit: Answering your questions
Recently, 10x Genomics launched its Chromium Nuclei Isolation Kit, which allows for easy and efficient sample prep from frozen tissues for single nuclei sequencing. The Nuclei Isolation Kit offers a unified sample prep workflow optimized for use with Chromium gene expression and chromatin accessibility assays, including Chromium Single Cell 3’ Gene Expression, 5’ Gene Expression, ATAC, Multiome ATAC + Gene Expression, and Single Cell Gene Expression Flex.
In an interactive webinar introducing the kit, 10x Genomics scientists Zuleyma Peralta, PhD, Product Manager, Sample Prep, and Mike Gibbons, PhD, Senior Scientist, Applications, discussed sample prep challenges and the ways that the kit can prep you for success. They fielded a variety of questions at the end. We share highlights from that live Q&A in this blog—keep reading to find answers to the most common questions, then watch the on-demand webinar for more details.
Sample compatibility across a wide range of tissue types
Have you tried this kit with my sample?
We’ve tested a wide range of human and mouse tissues. These include both easy-to-dissociate tissues like liver and fibrous, hard-to-dissociate tissues like heart, skeletal muscle, and skin. The specific tissues tested were chosen to reflect the wide range of possible input tissues expected to be used with this kit. A list of tissues tested is available here.
We expect this kit to be compatible across most mammalian tissues.
If your tissue is not currently listed, there may be other tissues we’ve tested with similar characteristics, which could be a good indicator that your sample is compatible. Users also asked about compatibility with fresh tissues and OCT-embedded tissues. We demonstrated compatibility with both!
We will continue to update the list of compatible tissues periodically as we test new samples.
Do I need to optimize lysis conditions for each tissue type?
All of the compatible tissues listed were run with the protocol straight out of the box, not necessitating any sample prep optimization, something usually needed with other nuclei isolation protocols/methods. If your sample is not in the list of tissues tested, we recommend running the workflow as described in the User Guide before attempting any further optimization.
The fact that no sample prep optimization is needed is especially important for users who may be working with precious clinical samples that cannot be used up for sample prep optimization and need to work on the first try. Additionally, if your lab works with multiple tissues, you simplify your nuclei isolation workflows by using one kit for all your nuclei isolation needs.
Streamlined sample prep workflow includes cleanup
Do I need to sort after using the Nuclei Isolation Kit?
Existing nuclei isolation protocols will require additional cleanup methods post-nuclei isolation to remove cellular debris and ambient RNA contaminants. These cleanup methods can require additional reagents, instrumentation (i.e., flow sorting), time, and handling, which can compromise the quality of the sample. With the Nuclei Isolation Kit, cleanup steps are incorporated into the workflow, meaning you get a clean nuclei suspension in one hour. We showed the example below, which demonstrates the efficacy of debris cleanup of the Chromium nuclei isolation workflow.
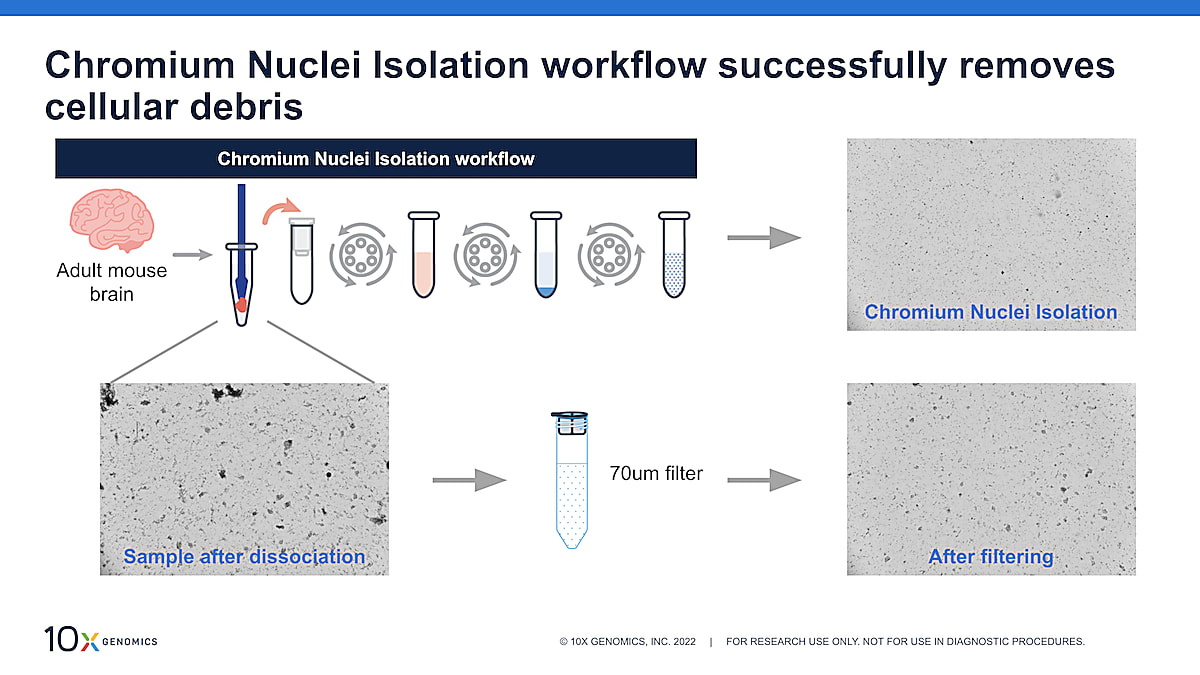
In Figure 1, we started off with adult mouse brain, a sample tissue type known to contain lots of debris like myelin, and dissociated that using a pestle and the Nuclei Isolation Kit lysis buffer (the first step of the nuclei isolation workflow). The image on the left shows the sample after dissociation. Lots of debris is present, including large pieces of debris. When we take that sample through the entire Nuclei Isolation Kit workflow, you can see that a lot of the debris is removed, leaving a clean nuclei suspension (top right). In comparison, if you were just to filter the nuclei suspension, some of the large debris is removed, but a lot make it through the filter (bottom right).
QC tips and tricks for optimizing yield and quality
How are the nuclei QC’d post isolation?
After isolation, the nuclei are counted using a fluorescent nucleic acid dye, such as AO/PI or ethidium homodimer, and a fluorescent-capable counter. Here, we evaluate nuclei yield and suspension quality (clumps, debris). We have observed that usage of a fluorescent nucleic dye improves nuclei count accuracy, as this only stains true nuclei. Though Trypan Blue is often used for counting, it can stain cellular debris and lead to increased nuclei estimates and underloading of the Chromium assays.
Because the Nuclei Isolation Kit workflow is quick, sample quality is maintained and high magnification imaging of nuclear membranes is not required (something we currently recommend for other protocols, as they tend to be lengthy with multiple sample handling steps).
How do I QC my tissue to check its quality before isolating nuclei using this kit?
RNA Integrity Number (RIN) can be used to assess tissue quality. This involves taking a small piece of your tissue, isolating RNA, and assessing the presence of ribosomal RNA peaks on a bioanalyzer instrument. A low RIN score indicates that the RNA within the sample may be degraded and can help you exclude poor quality samples. Keep in mind that RIN score evaluation is a bulk assessment and may not inform you on any cell stress your tissue may have undergone after collection that may show up in your final data or on ambient RNA/debris contaminants present within your sample.
For more information about the Nuclei Isolation Kit, visit the web page. And watch our on-demand webinar introducing the technology.
Resources:
