Visium CytAssist, Mouse Embryo, 11 mm Capture Area (FFPE)
Spatial Gene Expression dataset analyzed using Space Ranger 2.1.0
Learn about Visium Analysis
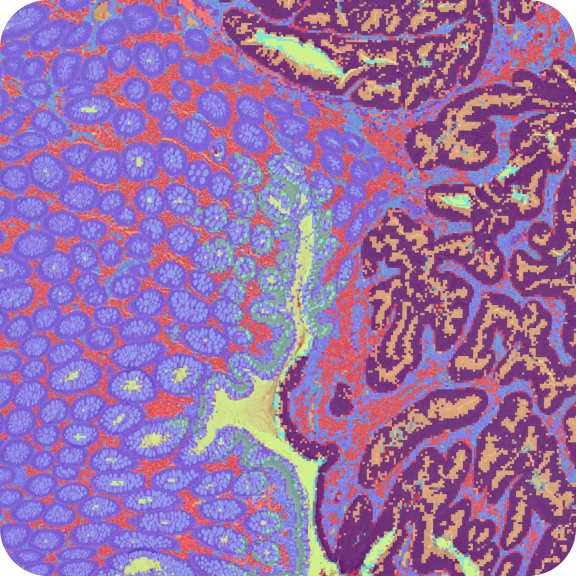
10x Genomics obtained FFPE Mouse Embryo tissue blocks from Avaden Biosciences. The tissue was sectioned as described in Visium CytAssist Spatial Gene Expression for FFPE – Tissue Preparation Guide Demonstrated Protocol CG000518. Tissue sections of 5 µm was placed on a standard glass slide, and H&E-stained following deparaffinization. Sections were coverslipped with 85% glycerol, imaged, decoverslipped, followed by dehydration & decrosslinking (Demonstrated Protocol CG000520). The glass slide with the tissue section was processed with the Visium CytAssist instrument to transfer analytes to a Visium CytAssist Spatial Gene Expression slide (11 mm Capture Area). The probe extension and library construction steps follow the standard Visium for FFPE workflow outside of the instrument.
The H&E image was acquired using an Olympus VS200 Slide Scanning Microscope with the following settings:
- Olympus Objective magnification: 20x UPLXAPO Objective
- Numerical Aperture: 0.8
- ScopeLED light source: Xcite Novum
- Camera: VS-264C
- Exposure: 410 µs
Libraries were prepared following the Visium CytAssist Spatial Gene Expression Reagent Kits for FFPE User Guide (CG000495).
- Sequencing instrument: Illumina NovaSeq, flow cell HNT5WDSX3 (lane 1-4)
- Sequencing configuration: 28bp read 1 (16bp Visium spatial barcode, 12bp UMI), 90bp read 2 (transcript), 10bp i7 sample barcode and 10bp i5 sample barcode
- Dual-Index set: SI-TS-D6
- Slide: V52Y09-019
- Area: B
This dataset is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) license. 10x citation guidelines available here.
