In this tutorial, you will learn how to run the spaceranger count pipeline on Visium Spatial Gene Expression data derived from a fresh frozen (FF) mouse brain coronal section.
To successfully run this tutorial, you should:
- Be comfortable in the Linux environment
- Have familiarity running command line tools
- Choose a compute platform
- Have access to a system that meets the minimum system requirements
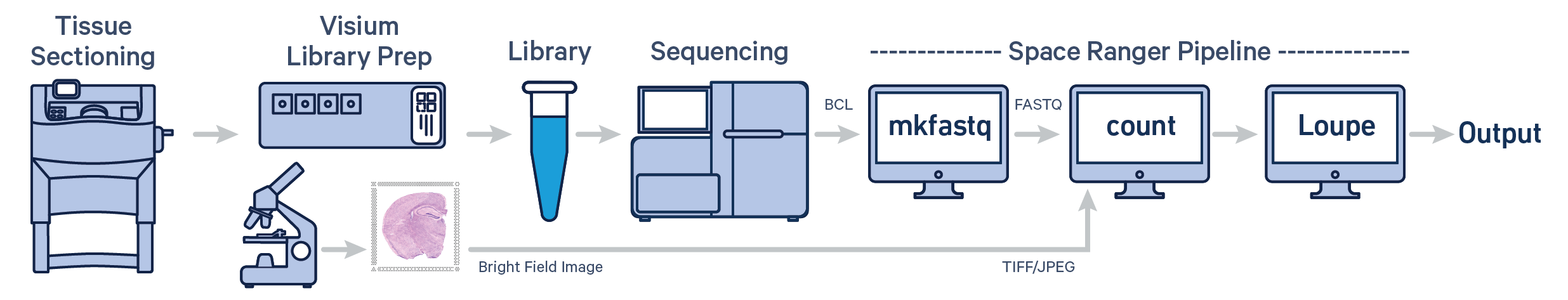
Visium Spatial Gene Expression data from FF tissues are analyzed using the spaceranger count pipeline. The pipeline inputs a microscope image of the Visium slide (in TIFF or JPEG format), a reference, and FASTQ files, and performs alignment, tissue and fiducial detection, and barcode/UMI counting. Outputs include the feature-spot matrices, clustering and differential gene expression (DGE) which can be further analyzed and visualized in Loupe Browser.
In this tutorial, you will analyze a mouse brain coronal section public dataset.
Key dataset features include:
- Tissue section of 10 µm thickness
- H&E image acquired using a Nikon Ti2-E microscope
- Sequencing Depth: 115,569 read pairs per spot
- Sequencing Coverage: Read 1 - 28 bp; Read 2 - 120 bp (transcript); i7 sample index - 10 bp; i5 sample index - 10 bp
- Visium Slide: V19L01-041
- Capture Area: C1
The following commands will be run in the working directory (spaceranger_tutorial) that was used to install Space Ranger on a compatible compute platform.
Both the raw sequencing files in FASTQ format, and the image in TIFF format, are available for download on the dataset page. For better organization, we will create a datasets folder prior to downloading the required file.
Download with curl command:
# Create datasets folder
mkdir datasets
# Download FASTQ to datasets folder
curl https://s3-us-west-2.amazonaws.com/10x.files/samples/spatial-exp/1.1.0/V1_Adult_Mouse_Brain/V1_Adult_Mouse_Brain_fastqs.tar -o datasets/V1_Adult_Mouse_Brain_fastqs.tar
# Download image file to datasets folder
curl https://cf.10xgenomics.com/samples/spatial-exp/1.1.0/V1_Adult_Mouse_Brain/V1_Adult_Mouse_Brain_image.tif -o datasets/V1_Adult_Mouse_Brain_image.tif
# Expected output
% Total % Received % Xferd Average Speed Time Time Time Current
Dload Upload Total Spent Left Speed
0 26.9G 0 135M 0 0 34.4M 0 0:13:22 0:00:03 0:13:19 34.4M
Alternatively, download with the wget command:
# Create datasets folder
mkdir datasets
# Download FASTQ to datasets folder
wget -P datasets/ https://s3-us-west-2.amazonaws.com/10x.files/samples/spatial-exp/1.1.0/V1_Adult_Mouse_Brain/V1_Adult_Mouse_Brain_fastqs.tar
# Download image file to datasets folder
wget -P datasets/ https://cf.10xgenomics.com/samples/spatial-exp/1.1.0/V1_Adult_Mouse_Brain/V1_Adult_Mouse_Brain_image.tif
# Expected output
Resolving s3-us-west-2.amazonaws.com (s3-us-west-2.amazonaws.com)... 52.218.217.16
Connecting to s3-us-west-2.amazonaws.com (s3-us-west-2.amazonaws.com)|52.218.217.16|:443...
connected.
HTTP request sent, awaiting response... 200 OK
Length: 28987985920 (27G) [application/x-tar]
Saving to: ‘V1_Adult_Mouse_Brain_fastqs.tar’
10% [=======> ] 3,179,419,763 36.2MB/s eta 11m 35s
Reference data
Download the latest version of the mouse transcriptome reference available from the Downloads page.
# Download mouse reference
curl -O https://cf.10xgenomics.com/supp/spatial-exp/refdata-gex-mm10-2020-A.tar.gz
# Expected output
% Total % Received % Xferd Average Speed Time Time Time Current
Dload Upload Total Spent Left Speed
1 9835M 1 158M 0 0 34.1M 0 0:04:48 0:00:04 0:04:44 34.1M
Extract files
After downloading the required files, the contents of the tar files need to be extracted.
# Extract sample FASTQ files
tar -xvf datasets/V1_Adult_Mouse_Brain_fastqs.tar -C datasets/ && rm datasets/V1_Adult_Mouse_Brain_fastqs.tar
# Extract mouse reference transcriptome
tar -xzvf refdata-gex-mm10-2020-A.tar.gz && rm refdata-gex-mm10-2020-A.tar.gz
# Expected output
# Sample FASTQ files
V1_Adult_Mouse_Brain_fastqs/
V1_Adult_Mouse_Brain_fastqs/V1_Adult_Mouse_Brain_S5_L002_I2_001.fastq.gz
V1_Adult_Mouse_Brain_fastqs/V1_Adult_Mouse_Brain_S5_L001_R1_001.fastq.gz
...
# Reference mouse transcriptome
refdata-gex-mm10-2020-A/
refdata-gex-mm10-2020-A/fasta/
refdata-gex-mm10-2020-A/fasta/genome.fa
...
This will create two additional folders (highlighted in yellow) within the working directory.
1spaceranger_tutorial 2├── datasets 3│ ├── V1_Adult_Mouse_Brain_fastqs 4│ └── V1_Adult_Mouse_Brain_image.tif 5├── refdata-gex-mm10-2020-A 6└── spaceranger-2.0.0
You can now build the spaceranger count command to run from your working directory (spaceranger_tutorial). If running from a different directory, amend the paths accordingly to avoid any errors.
spaceranger count --id="V1_Adult_Mouse_Brain" \
--transcriptome=refdata-gex-mm10-2020-A \
--fastqs=datasets/V1_Adult_Mouse_Brain_fastqs \
--image=datasets/V1_Adult_Mouse_Brain_image.tif \
--slide=V19L01-041 \
--area=C1 \
--create-bam=false \
--localcores=16 \
--localmem=128
Below are brief descriptions of the command line options:
| Option | Description |
|---|---|
--id | The id must be a unique string and will be used to name the resulting folder with all of the pipeline outputs. |
--transcriptome | The path to the species specific pre-compiled transcriptome files. Note that you can either provide the relative path as shown above or the absolute path to this folder. As the tissue sample was of mouse origin, we provide the path to the mouse reference transcriptome refdata-gex-mm10-2020-A |
--fastqs | The path to the folder containing FASTQ files. The path can be relative as shown above or absolute. The relative path is /datasets/V1_Adult_Mouse_Brain_fastqs |
--image | The path to a single brightfield image with H&E staining in either TIFF or JPEG formats. |
--slide | The Visium slide serial number. |
--area | The Capture Area identifier on the Visium slide. It can be one of four values: A1, B1, C1 or D1. |
--localcores | The number of CPU cores available to run the spaceranger count pipeline. The maximum upper limit for your specific compute system is determined using the sitecheck subcommand. |
--localmem | The max memory in GB available to run the spaceranger count pipeline. The maximum upper limit for your specific compute system is determined using the sitecheck subcommand. |
At the start of the run, you should see the preflight checks printed to the command line.
# With internet access
# Run spaceranger count
spaceranger count --id="V1_Adult_Mouse_Brain" \
--description="Adult Mouse Brain (Coronal)" \
--transcriptome=refdata-gex-mm10-2020-A \
--fastqs=datasets/V1_Adult_Mouse_Brain_fastqs \
--image=datasets/V1_Adult_Mouse_Brain_image.tif \
--slide=V19L01-041 \
--area=C1 \
--localcores=16 \
--localmem=128
# Without internet access
spaceranger count --id="V1_Adult_Mouse_Brain" \
--description="Adult Mouse Brain (Coronal)" \
--transcriptome=refdata-gex-mm10-2020-A \
--fastqs=datasets/V1_Adult_Mouse_Brain_fastqs \
--image=datasets/V1_Adult_Mouse_Brain_image.tif \
--slide=V19L01-041 \
--slidefile=V19L01-041.gpr \
--area=C1 \
--localcores=16 \
--localmem=128
# Expected output
Martian Runtime - v4.0.5
Running preflight checks (please wait)...
Checking sample info...
Checking FASTQ folder...
Checking reference...
Checking reference_path...
Checking optional arguments...
...
Successful completion of the pipeline is indicated by a list of output files.
Outputs:
- Run summary HTML: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/web_summary.html
- Outputs of spatial pipeline:
aligned_fiducials: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/spatial/aligned_fiducials.jpg
detected_tissue_image: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/spatial/detected_tissue_image.jpg
scalefactors_json: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/spatial/scalefactors_json.json
tissue_hires_image: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/spatial/tissue_hires_image.png
tissue_lowres_image: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/spatial/tissue_lowres_image.png
cytassist_image: null
aligned_tissue_image: null
tissue_positions: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/spatial/tissue_positions.csv
spatial_enrichment: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/spatial/spatial_enrichment.csv
barcode_fluorescence_intensity: null
- Run summary CSV: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/metrics_summary.csv
- Correlation values between isotypes and Antibody features: null
- BAM: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/possorted_genome_bam.bam
- BAM BAI index: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/possorted_genome_bam.bam.bai
- BAM CSI index: null
- Filtered feature-barcode matrices MEX: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/filtered_feature_bc_matrix
- Filtered feature-barcode matrices HDF5: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/filtered_feature_bc_matrix.h5
- Unfiltered feature-barcode matrices MEX: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/raw_feature_bc_matrix
- Unfiltered feature-barcode matrices HDF5: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/raw_feature_bc_matrix.h5
- Secondary analysis output CSV: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/analysis
- Per-molecule read information: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/molecule_info.h5
- Loupe Browser file: /spaceranger_tutorial/V1_Adult_Mouse_Brain/outs/cloupe.cloupe
- Feature Reference: null
- Target Panel file: null
- Probe Set file: null
Waiting 6 seconds for UI to do final refresh.
Pipestance completed successfully!
After the run is completed, the working directory will have a new folder named V1_Adult_Mouse_Brain (which was provided to the --id argument) that contains all the metadata and outputs generated from the spaceranger count pipeline:
V1_Adult_Mouse_Brain
├── _cmdline
├── _filelist
├── _finalstate
├── _invocation
├── _jobmode
├── _log
├── _mrosource
├── outs
├── _perf
├── _sitecheck
├── SPATIAL_RNA_COUNTER_CS
├── _tags
├── _timestamp
├── _uuid
├── V1_Adult_Mouse_Brain.mri.tgz
├── _vdrkill
└── _versions
V1_Adult_Mouse_Brain.mri.tgzcontains diagnostic information helpful to 10x Genomics support to resolve any errors_sitecheckcaptures the system configuration, similar to the sitecheck subcommand_timestampcontains information on pipeline runtimes._cmdlinecaptures thecountcommand provided to run the pipeline_versionscontains both thespacerangerand Martian versions used in the run * Theoutsfolder contain all the calculated results.

You can further explore and understand these results by
- Browsing the web_summary.html file.
- Opening the
.cloupefile in Loupe Browser - Referring to the Understanding Outputs page to explore individual files