Starting your spatial transcriptomics journey strong

What is spatial transcriptomics?
Spatial transcriptomics measures both the expression of your genes of interest and where in the tissue they’re expressed. By doing so, it enables huge leaps in characterizing cell–cell interactions, signaling networks, cell tissue composition, and biomarker discovery/validation that other technologies simply can’t match. For example, a cancer researcher using spatial methods can not only quantify relative immune cell abundance in immunotherapy responders versus non-responders, but determine whether there’s an association between better responses and immune cells being positioned close to the tumor. Simply put, spatial transcriptomics adds a new dimension to gene expression analyses that other methods can’t match, increasing the impact of your work. Simply put, spatial transcriptomics adds a new dimension to gene expression analyses that other methods can’t match, increasing the impact of your work.
What can spatial techniques do for my research?
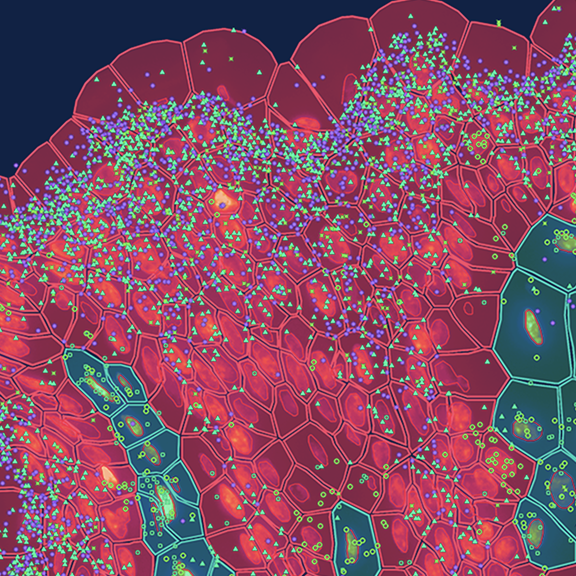

Identifying cell localization as a biomarker
Combining cell types with their locations in the tissue context can unlock a new kind of biomarker that legacy methods miss.
Spatial biomarkers of response to neoadjuvant therapy in muscle-invasive bladder cancer: the DUTRENEO trial
Exploring how local environments can influence cell state
Pathology, inflammation, and other tissue microenvironments can induce novel cell states, providing deeper insights into the mechanisms of disease.
An exhausted-like microglial population accumulates in aged and APOE4 genotype Alzheimer’s brains
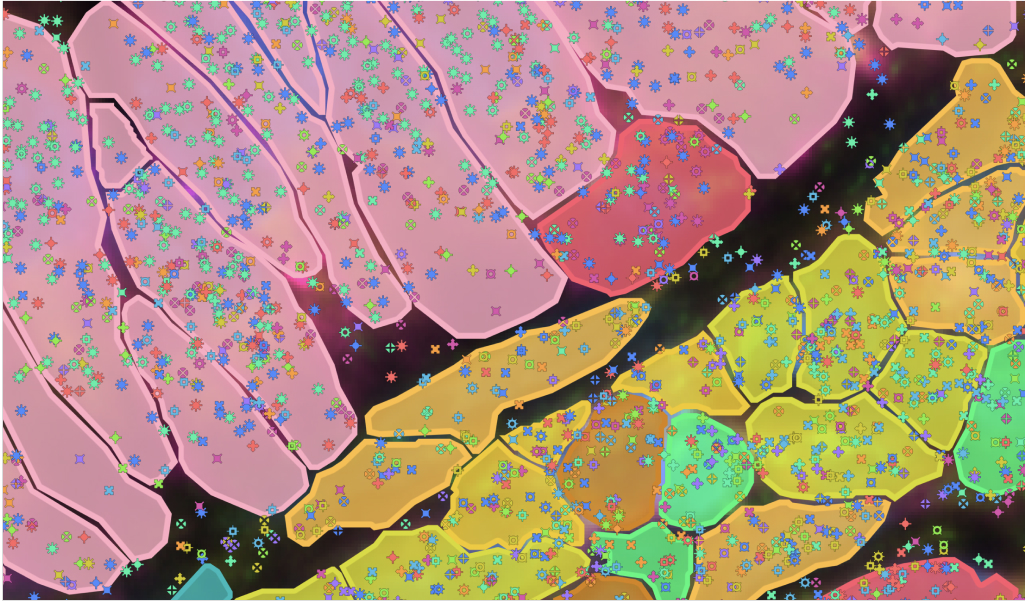

Untangling cell–cell interactions in the tumor microenvironment
Researchers used the Visium HD assay in human colorectal cancer FFPE samples to identify spatially distinct immune cell populations in and around the tumor. They then validated their findings, and identified an anti-tumor macrophage niche, with the Xenium platform.
High-definition spatial transcriptomic profiling of immune cell populations in colorectal cancer

Explore a new world of discovery with spatial biology
See how researchers around the world have used 10x Genomics Xenium and Visium assays for high-impact publications in research areas like oncology, neuroscience, immunology, and more.
What are the advantages of spatial?
| Spatial transcriptomics | Legacy ISH methods | IHC or IF | |
| Benefits | High-resolution single cell gene expression with native tissue context greatly expands discovery power. Preservation of spatial context of cells, empowering untangling of cell–cell interactions, characterization of cell neighborhoods, discovery of rare cells, and more. Some assays enable spatial multiomics for same-cell RNA and protein (as well as histology). | Relatively low-cost technique that uses common reagents. Spatial context for gene expression, down to single-molecule resolution. Spatial context for gene expression, down to single-molecule resolution | Affordable procedure that can be performed with few reagents. Tissue context of gene expression. Tissue context of gene expression |
| Challenges | Needs robust chemistry to avoid high background noise and elevated false discovery rates. Necessitates profiling analytes in the same cell or section (not sequential sections) to provide biologically accurate spatial multiomics. Can require specialized analysis; not all assays include integrated and/or recommended analysis tools. | Low-plex, no protein staining, and (depending on detection method) may have low resolution. Multi-step procedure that may require use of hazardous reagents, including radioactivity. Probes must be designed and optimized by the user. Multi-step procedure that may require use of hazardous reagents, including radioactivity Probes must be designed and optimized by the user | Specificity of antibodies can be variable (stains can be non-specific), often requiring antibody validation. Low-plex with results that make it challenging to quantify results. Expensive equipment needed as part of a multi-step procedure. May be challenging to quantify results Expensive equipment needed as part of a multi-step procedure |
How can spatial transcriptomics and multiomics uniquely benefit your research?
Learn how imaging- and sequencing-based spatial transcriptomics offers a whole new dimension of insights by letting you add tissue context to your single cell gene expression changes and layering protein analysis on top of RNA.
How can spatial enhance my research?
❌ Cutting-edge spatial transcriptomics is prohibitively expensive
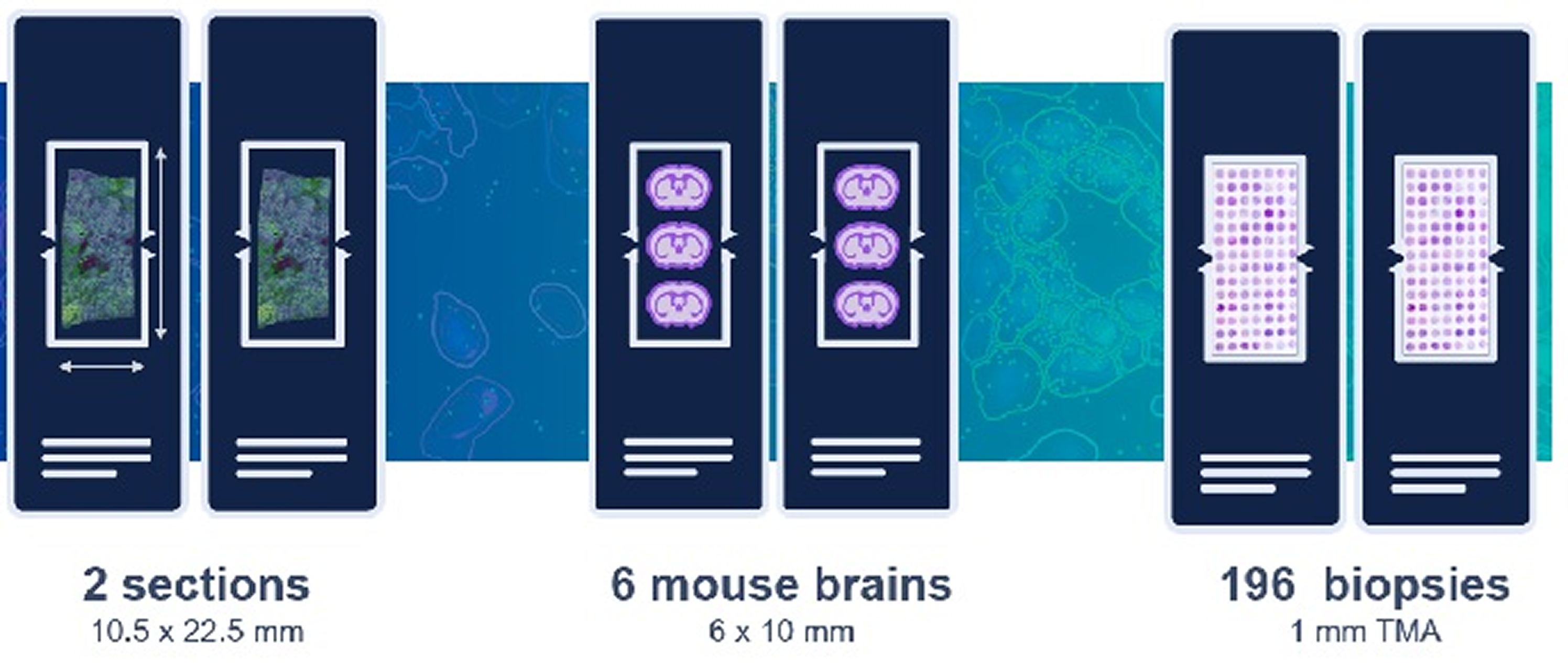
✅ With good experimental planning and tissue placement, a single Xenium run can provide data on multiple samples, making it remarkably cost-effective and providing higher-power pilot studies
Optimizing tissue placement on a single Xenium slide (see below) enables you to make the most of your analyses. Explore how researchers like Dr. Nick Banovich are achieving enormous per-sample cost savings with the Xenium platform.
❌ Spatial transcriptomics is a “nice-to-have,” acting as a supplement to your main experiment(s) of interest
✅ Spatial transcriptomics is a need-to-have for deeper biological insights
Only single cell spatial imaging technologies like the Xenium platform can analyze cellular neighborhoods, cell–cell interactions in native tissue context, cell proximity to pathology and disease-associated cell types, and more modalities, enabling you to capture information from your tissue that other technologies simply can’t match.
❌ You can’t identify high-homology transcripts, SNVs, and isoforms with spatial context
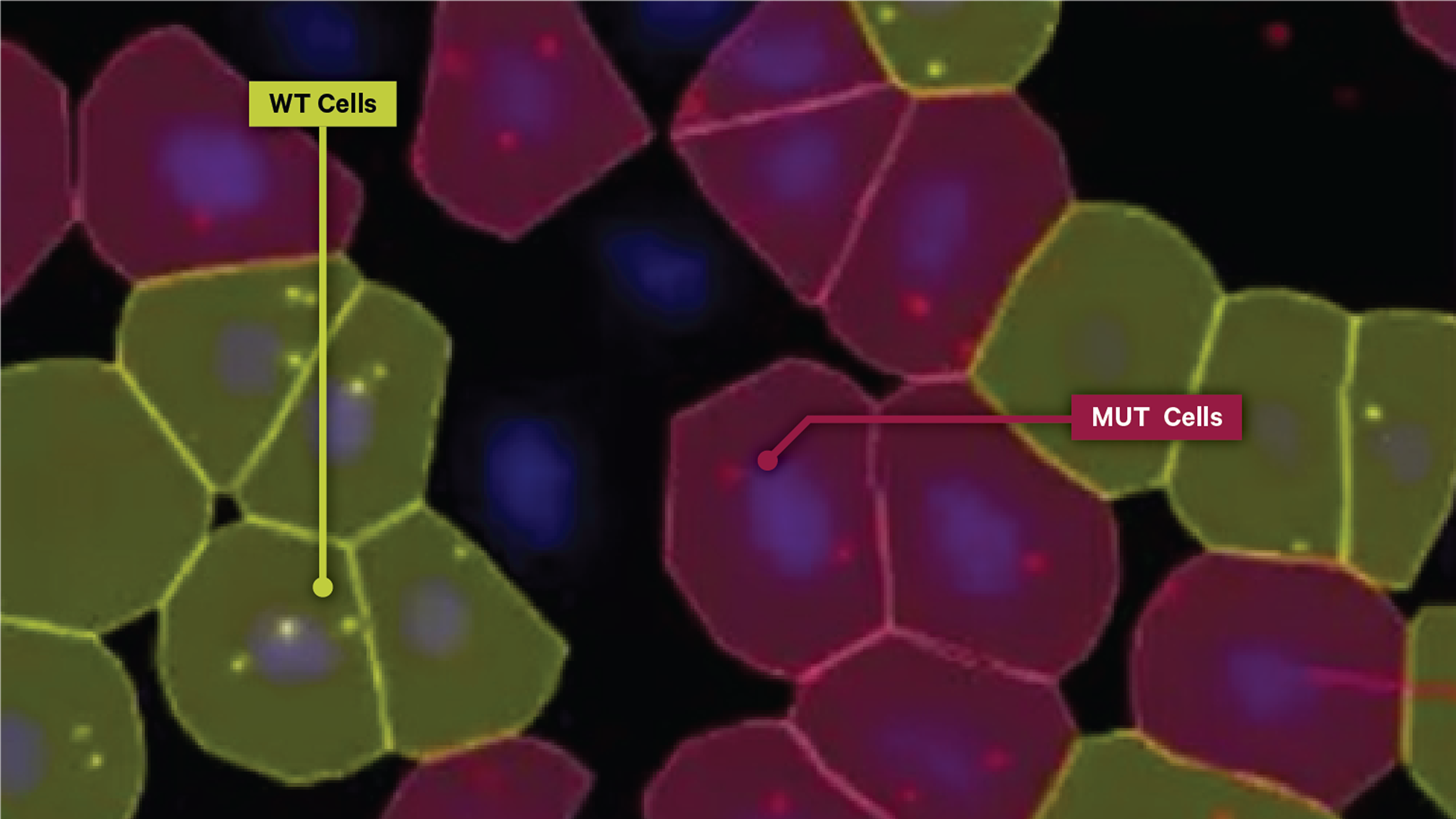
✅ The Xenium platform's unparalleled specificity and sensitivity enables you to discriminate between high-homology RNAs with spatial context at subcellular resolution
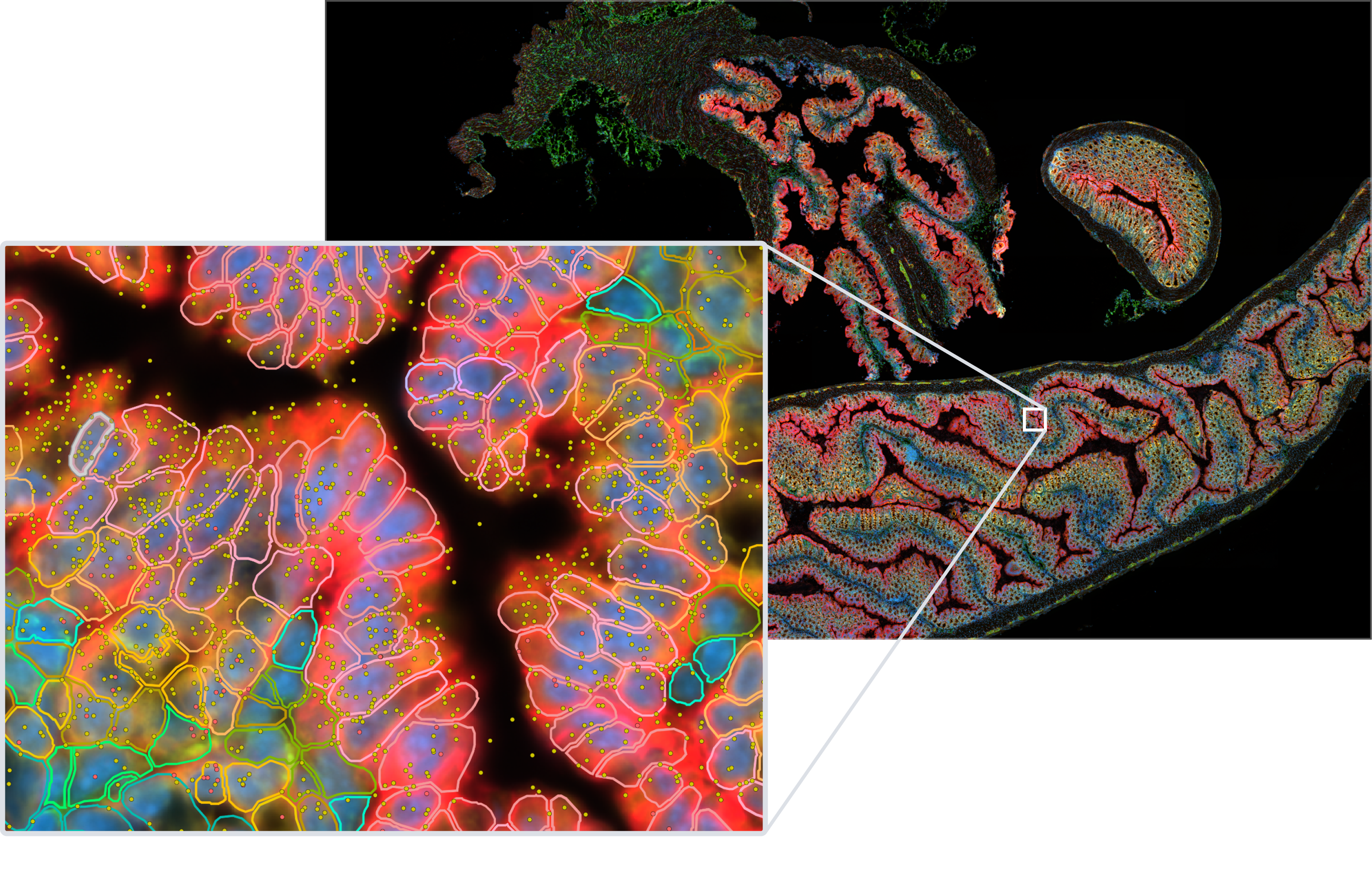
Capture a more complete picture of your biology’s transcriptomic architecture, such as identifying cancerous versus non-cancerous cells (see below), by teasing apart biologically relevant SNVs and isoforms in their native tissue context.
❌ Analyzing single cell spatial data is highly complex and requires bioinformatics expertise to unlock insights
✅ The Xenium platform makes single cell spatial insights easy with integrated, automated imaging and data analysis
Start with Xenium Onboard Analysis, which processes data in parallel with imaging steps, to get interpretation-ready data the moment your run is done. Then leverage our intuitive and integrated software suite to easily characterize cell types, incorporate cell labels from scRNA-seq datasets, map cell pathways, plot tissue networks, and more for deeper insights into your biology of interest.
Why choose 10x for your spatial research?

Xenium
Single cell spatial imaging with subcellular resolution
Robust single cell spatial discovery of up to 5,000 genes at subcellular resolution
Same-section spatial multiomics for analysis of RNA, protein, and histology
Maximum experimental flexibility with customizable pre-designed and fully custom panels
Visium
NGS-based spatial transcriptomics at single cell scale
Expanded discovery with whole transcriptome spatial gene expression at single cell scale
Can be used with pre-sectioned/archival slides
Compatible with non-human and mouse species

Compare assays
Compare spatial assays by compatible species and samples, multiomic capabilities, run configuration, resolution, multiplexing options, compatible instruments, and more.