Single Cell Analysis
A whole new visual canvas to explore single cell data
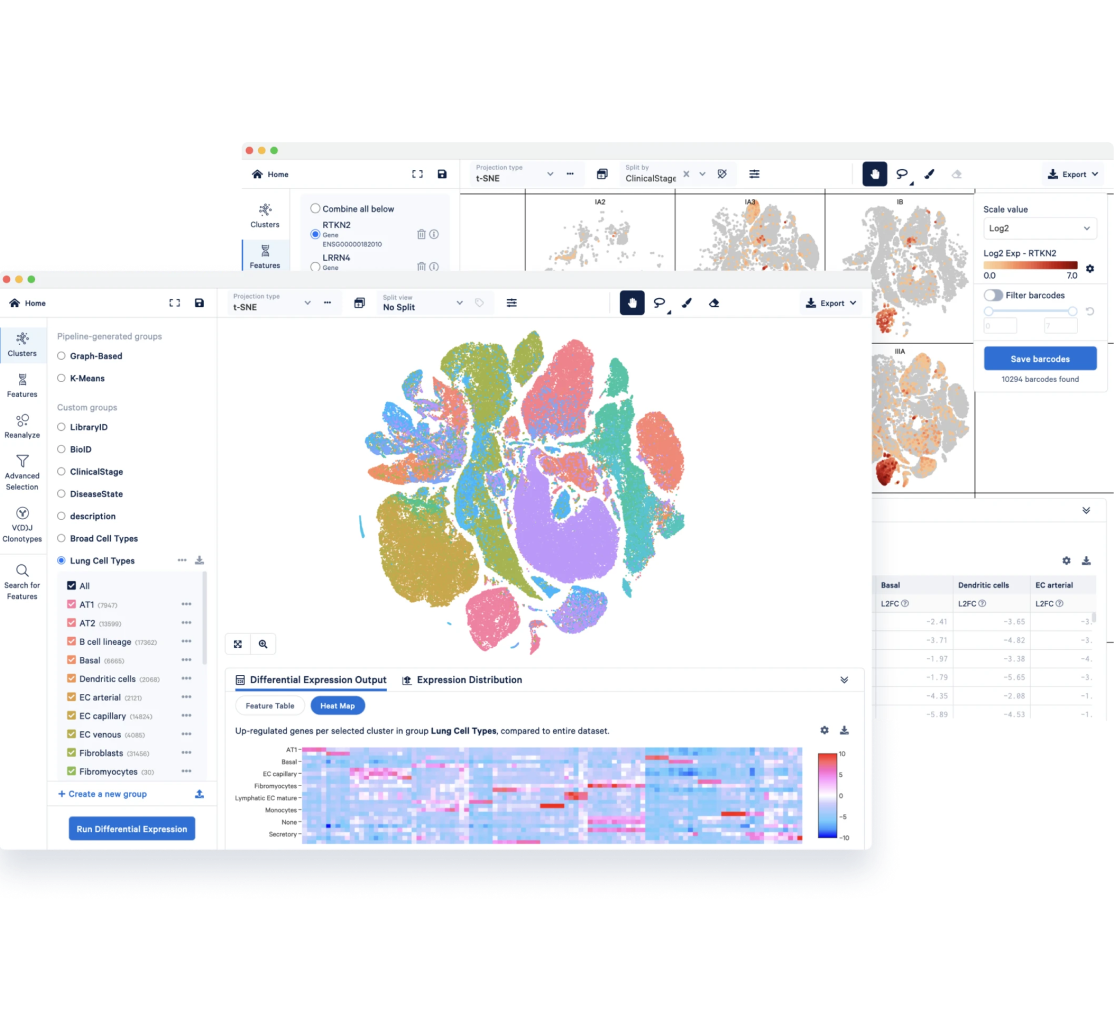
Your roadmap to single cell data analysis

Cell Ranger analysis pipelines
Transform raw sequences into single cell insights
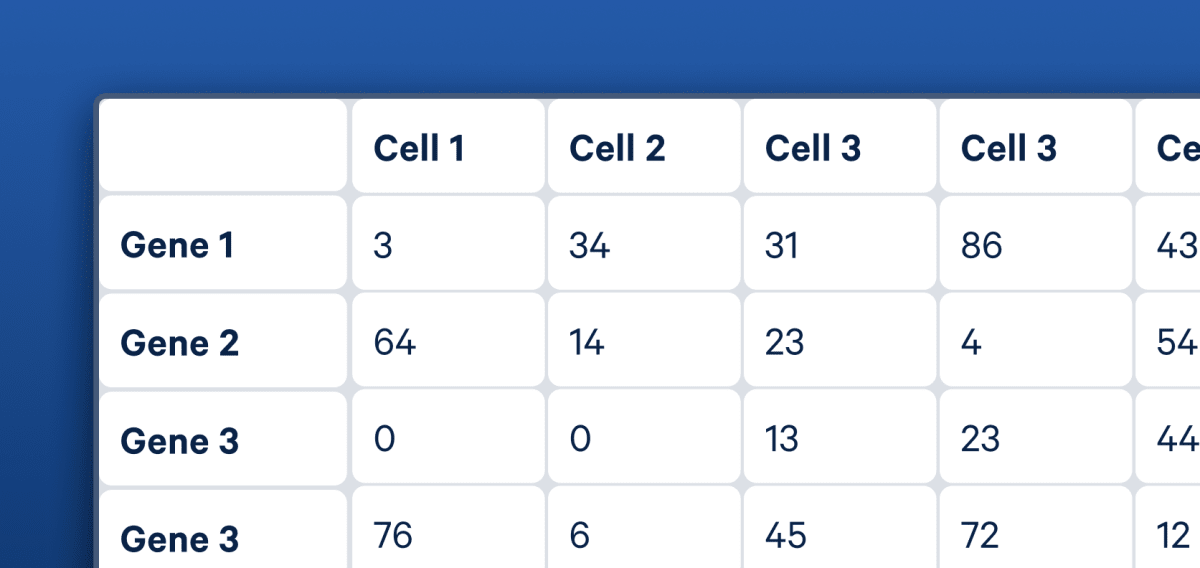
Start with Cell Ranger to align reads, generate cell-feature matrices, perform clustering and secondary analysis
Cell Ranger can process raw sequencing data, perform QC, identify cell barcodes, and quantify gene expression levels at the single cell level.
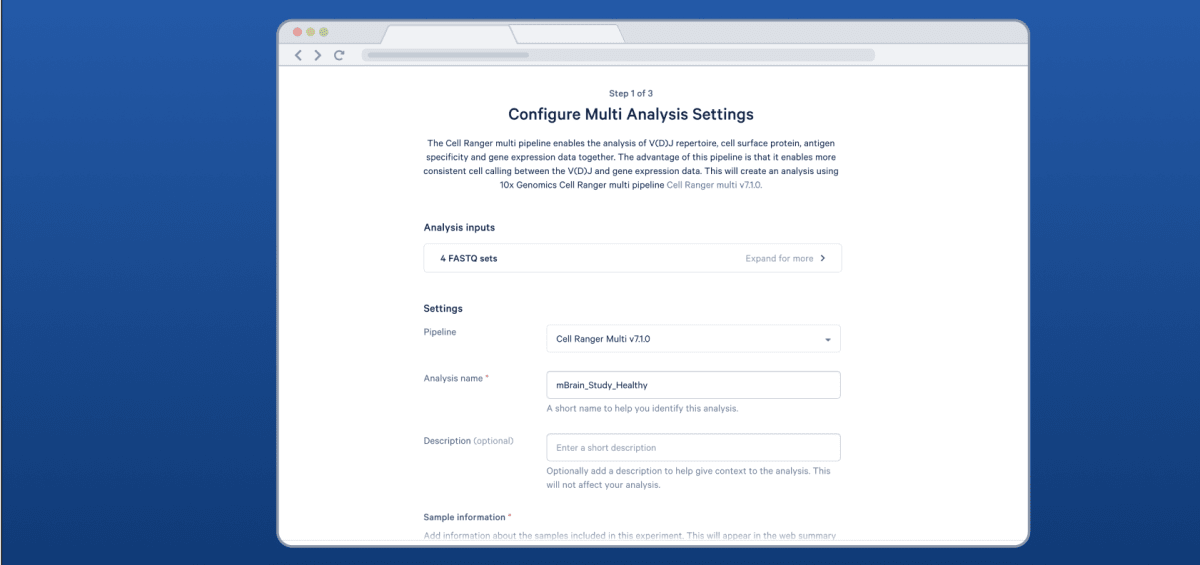
Accelerate your time to results with Cloud Analysis, a web app for running Cell Ranger
Access Cell Ranger from your web browser with Cloud Analysis.
Loupe Browser
10 ways to dig into your data with Loupe Browser
Use Loupe to dive deeper into your Seurat analysis, no code required
LoupeR is an R package that creates a Loupe Browser file from a Seurat object. With LoupeR, seamlessly explore Seurat data in Loupe without command-line data wrangling.Install LoupeRFind differentially expressed genes in treatment vs. control experiments
Easily perform and iterate on multi-sample comparisons.Rapidly visualize expression of your favorite genes
Quickly plot your differential expression results
Explore interactive heat maps and violin plots of differentially expressed genes.Easily annotate cell types
Identify and annotate broad and fine-grained human cell types with Automated Cell Annotation.Generate fast and interactive differential expression analyses
Reanalyze subsets of data to find new cell types
Narrow in on your cell types of interest or filter out low quality data.Create high-resolution figures in a few clicks
Loupe Browser makes it easy to generate SVG and PNG formatted images to share in presentations or publications.Characterize cells using advanced filters
Hone in on particular sets of barcodes by creating a custom set of rules with boolean operators.Explore many modalities of data, including single cell V(D)J, ATAC, and spatial gene expression
Loupe Browser supports the full suite of 10x Genomics single cell and spatial datasets.
Explore single cell data
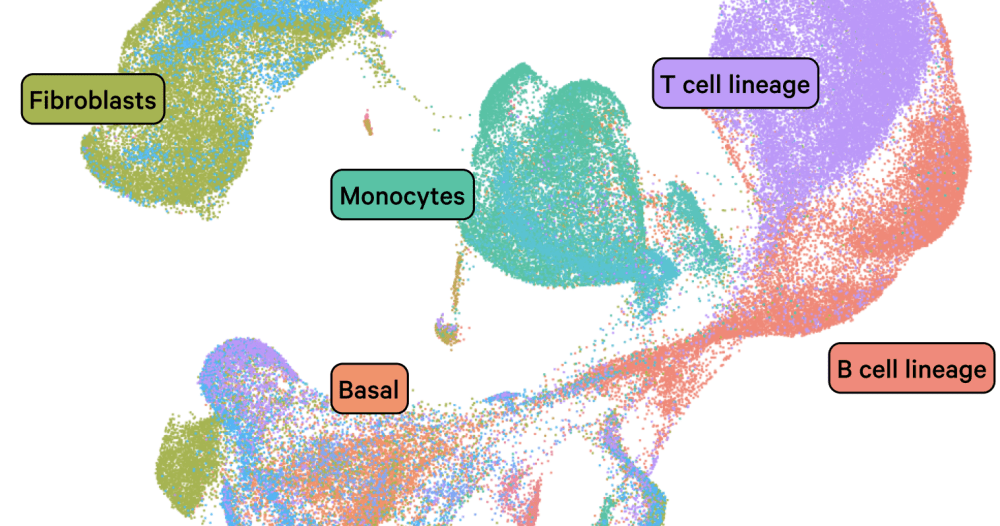
Aggregate of ~900k Human Non Small Cell Lung Cancer and Normal Adjacent Cells
Species
HumanTissue
LungNumber of cells
897,733Assay used
Single Cell Gene Expression Flex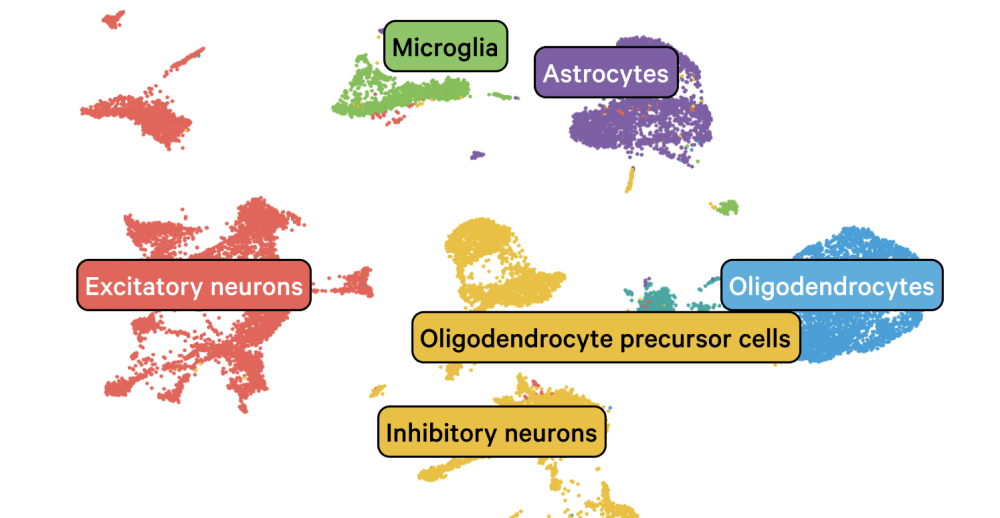
Alzheimer's Disease Mouse Model - Brain Coronal Sections from One Hemisphere Over a Time Course
Species
MouseTissue
BrainNumber of cells
33,459